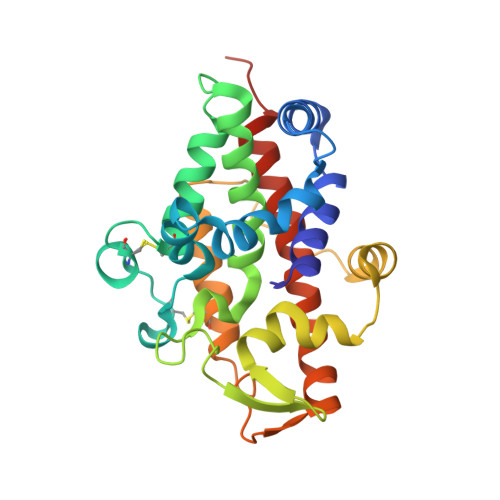
Substrate preference, RNA binding and active site versatility of Stenotrophomonas maltophilia nuclease SmNuc1, explained by a structural study.
Adamkova, K., Trundova, M., Koval, T., Hustakova, B., Kolenko, P., Duskova, J., Skalova, T., Dohnalek, J.(2025) FEBS J 292: 129-152
- PubMed: 39361520 Search on PubMed
- DOI: https://doi.org/10.1111/febs.17265
- Primary Citation Related Structures:
8QJL, 8QJM, 8QJN, 8QJO, 8QJP, 8QJQ, 9EMG - PubMed Abstract:
Nucleases of the S1/P1 family have important applications in biotechnology and molecular biology. We have performed structural analyses of SmNuc1 nuclease from Stenotrophomonas maltophilia, including RNA cleavage product binding and mutagenesis in a newly discovered flexible Arg74-motif, involved in substrate binding and product release and likely contributing to the high catalytic rate. The Arg74Gln mutation shifts substrate preference towards RNA. Purine nucleotide binding differs compared to pyrimidines, confirming the plasticity of the active site. The enzyme-product interactions indicate a gradual, stepwise product release. The activity of SmNuc1 towards c-di-GMP in crystal resulted in a distinguished complex with the emerging product 5'-GMP. This enzyme from an opportunistic pathogen relies on specific architecture enabling high performance under broad conditions, attractive for biotechnologies.
- Institute of Biotechnology, Czech Academy of Sciences, Vestec, Czech Republic.
Organizational Affiliation: