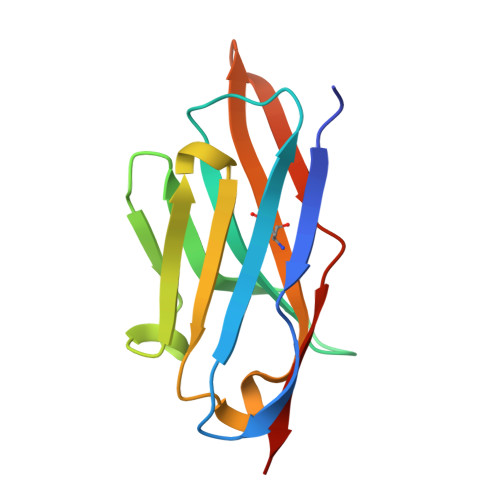
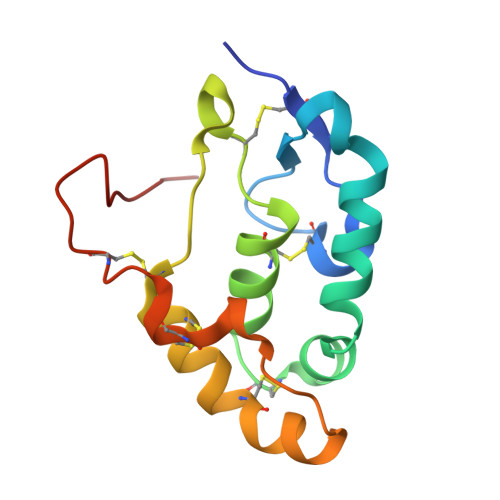
Structural insights into Frizzled3 through nanobody modulators.
Hillier, J., Zhao, Y., Carrique, L., Malinauskas, T., Ruza, R.R., Chang, T.H., Yi, G., Duyvesteyn, H.M.E., Yu, J., Lu, W., Pardon, E., Steyaert, J., Zhu, Y., Ni, T., Jones, E.Y.(2024) Nat Commun 15: 7228-7228
- PubMed: 39174501 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-51451-1
- Primary Citation Related Structures:
8Q7O, 8QW4 - PubMed Abstract:
The Wnt receptor Frizzled3 (FZD3) is important for brain axonal development and cancer progression. We report structures of FZD3 in complex with extracellular and intracellular binding nanobodies (Nb). The crystal structure of Nb8 in complex with the FZD3 cysteine-rich domain (CRD) reveals that the nanobody binds at the base of the lipid-binding groove and can compete with Wnt5a. Nb8 fused with the Dickkopf-1 C-terminal domain behaves as a FZD3-specific Wnt surrogate, activating β-catenin signalling. The cryo-EM structure of FZD3 in complex with Nb9 reveals partially resolved density for the CRD, which exhibits positional flexibility, and a transmembrane conformation that resembles active GPCRs. Nb9 binds to the cytoplasmic region of FZD3 at the putative Dishevelled (DVL) or G protein-binding site, competes with DVL binding, and inhibits GαS coupling. In combination, our FZD3 structures with nanobody modulators map extracellular and intracellular interaction surfaces of functional, and potentially therapeutic, relevance.
- Division of Structural Biology, Wellcome Centre for Human Genetics, University of Oxford, Oxford, UK.
Organizational Affiliation: