HIF prolyl hydroxylase 2 in complex with HIF2alpha-CODD
Fiorini, G., Figg Jr, W.D., Schofield, C.J.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Egl nine homolog 1 | 233 | Homo sapiens | Mutation(s): 0 Gene Names: EGLN1, C1orf12, PNAS-118, PNAS-137 EC: 1.14.11.29 |  | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9GZT9 GTEx: ENSG00000135766 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9GZT9 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Endothelial PAS domain-containing protein 1 | 20 | Homo sapiens | Mutation(s): 0 |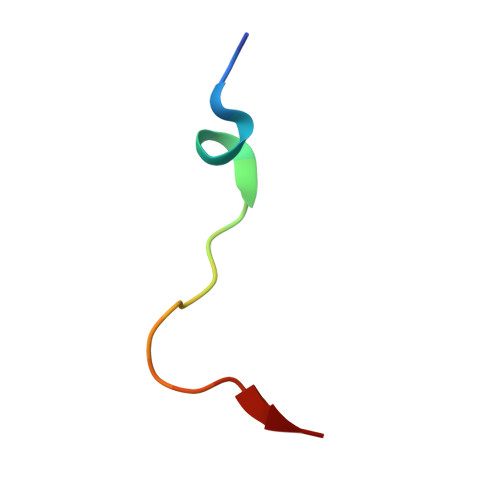  | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q99814 GTEx: ENSG00000116016 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q99814 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ACT Download:Ideal Coordinates CCD File | C [auth A], D [auth A], E [auth A] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 37.888 | α = 90 |
| b = 42.824 | β = 90 |
| c = 130.807 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| DIALS | data scaling |
| xia2 | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Wellcome Trust | United Kingdom | -- |
| University of Oxford, Medical Sciences Internal Fund (MSIF) | United Kingdom | -- |
| Cancer Research UK | United Kingdom | -- |
| Biotechnology and Biological Sciences Research Council (BBSRC) | United Kingdom | -- |