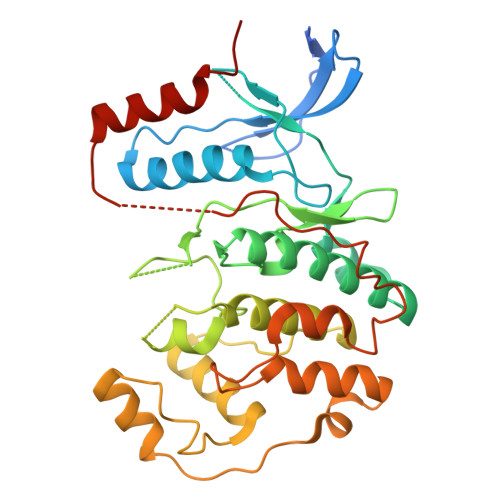
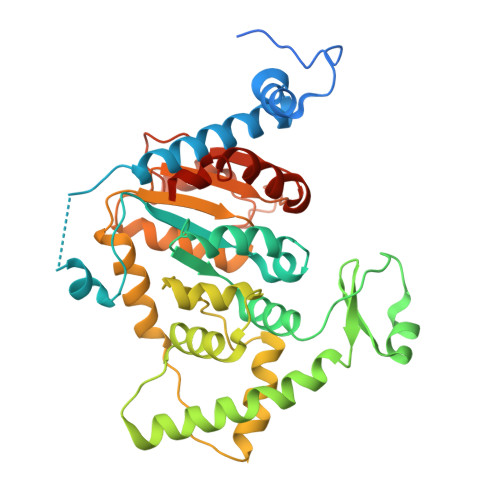
ERK1/2 interaction with DHPS regulates eIF5A deoxyhypusination independently of ERK kinase activity.
Becker, A.E., Kochanowski, P., Wu, P.K., Wator, E., Chen, W., Guchhait, K., Biela, A.P., Grudnik, P., Park, J.I.(2024) Cell Rep 43: 114831-114831
- PubMed: 39392755 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2024.114831
- Primary Citation Related Structures:
8PVU - PubMed Abstract:
This study explores a non-kinase effect of extracellular regulated kinases 1/2 (ERK1/2) on the interaction between deoxyhypusine synthase (DHPS) and its substrate, eukaryotic translation initiation factor 5A (eIF5A). We report that Raf/MEK/ERK activation decreases the DHPS-ERK1/2 interaction while increasing DHPS-eIF5A association in cells. We determined the cryoelectron microscopy (cryo-EM) structure of the DHPS-ERK2 complex at 3.5 Å to show that ERK2 hinders substrate entrance to the DHPS active site, subsequently inhibiting deoxyhypusination in vitro. In cells, impairing the ERK2 activation loop, but not the catalytic site, prolongs the DHPS-ERK2 interaction irrespective of Raf/MEK signaling. The ERK2 Ser-Pro-Ser motif, but not the common docking or F-site recognition sites, also regulates this complex. These data suggest that ERK1/2 dynamically regulate the DHPS-eIF5A interaction in response to Raf/MEK activity, regardless of its kinase function. In contrast, ERK1/2 kinase activity is necessary to regulate the expression of DHPS and eIF5A. These findings highlight an ERK1/2-mediated dual kinase-dependent and -independent regulation of deoxyhypusination.
- Department of Biochemistry, Medical College of Wisconsin, Milwaukee, WI 53226, USA.
Organizational Affiliation: