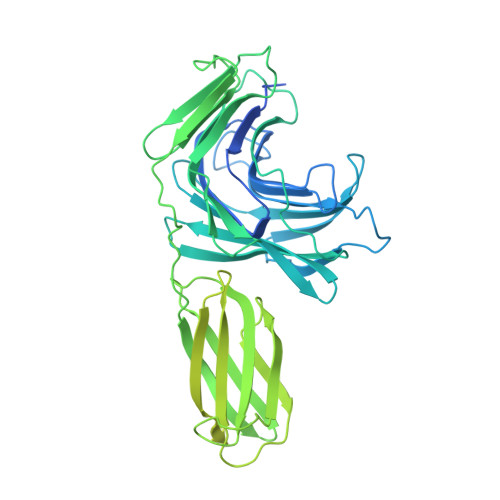
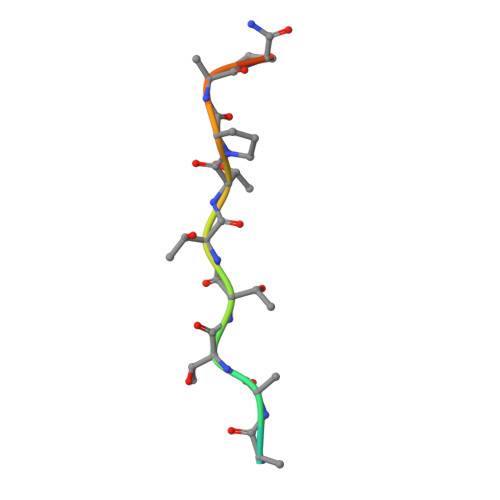
A family of di-glutamate mucin-degrading enzymes that bridges glycan hydrolases and peptidases
Narimatsu, Y., Bull, C., Taleb, V., Liao, Q., Companon, I., Sanchez-Navarro, D., Durbesson, F., Vincentelli, R., Hansen, L., Corzana, F., Rovira, C., Henrissat, B., Clausen, H., Joshi, H.J., Hurtado-Guerrero, R.(2024) Nat Catal