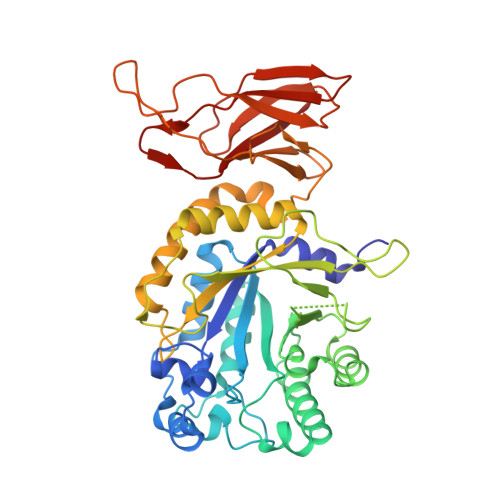
Unveiling the structural bases of alpha-L-fucosidase B activity through mutants boosting transfucosylation efficiency.
Becerra, J.E., Gallego Del Sol, F., Rubio-Del-Campo, A., Rodriguez-Diaz, J., Monedero, V., Marina, A., Yebra, M.J.(2025) Int J Biol Macromol 311: 143462-143462
- PubMed: 40286956 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2025.143462
- Primary Citation Related Structures:
8OZT, 8OZU, 9HY7, 9HYJ, 9HYX, 9HZ1 - PubMed Abstract:
The AlfB α-L-fucosidase from Lacticaseibacillus paracasei exhibits high specificity on fucosyl-α1,3-N-acetylglucosamine, achieving yields of 30 % in transfucosylation reactions for its synthesis. By random mutagenesis we selected AlfB variants with enhanced transfucosylation activity. Expression of a collection of alfB mutants in E. coli resulted in the isolation of eighteen clones with reduced activity on p-nitrophenyl-α-L-fucopyranoside. The AlfB variants carried diverse amino substitutions, leading to modifications in their enzymatic parameters. In some cases, these changes increased transfucosylation yields, although no direct correlation was observed between k cat or K m values and the yields. One particular AlfB mutant (M58) achieved 100 % yield in the synthesis of fucosyl-α1,3-N-acetylglucosamine. This enzyme contained three amino acid substitutions (N196S, V261M and N346K); however, further analysis confirmed that the N346K mutation was sufficient to generate the maximum yield. Elucidation of the tridimensional structure of AlfB and AlfBM58 through X-ray crystallography allowed us to propose a mechanism by which the mutation at position 346, located in a loop close to the active site of an adjacent monomer in the protein tetramer, enhanced transfucosylation over hydrolysis of fucosyl-α1,3-N-acetylglucosamine. This study paves the way for designing novel AlfB variants as tools for the efficient enzymatic synthesis of regio-specific fucosyl-oligosaccharides of biotechnological interest.
- Laboratorio de Bacterias Lácticas y Probióticos, Departamento de Biotecnología de Alimentos, Instituto de Agroquímica y Tecnología de Alimentos (IATA-CSIC), Valencia, Spain.
Organizational Affiliation: