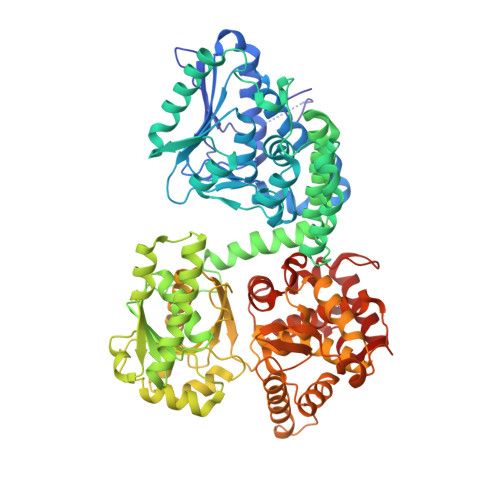
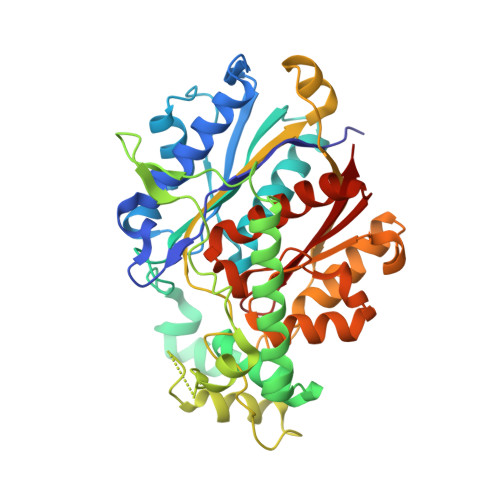
Crystallographic fragment-binding studies of the Mycobacterium tuberculosis trifunctional enzyme suggest binding pockets for the tails of the acyl-CoA substrates at its active sites and a potential substrate-channeling path between them.
Dalwani, S., Metz, A., Huschmann, F.U., Weiss, M.S., Wierenga, R.K., Venkatesan, R.(2024) Acta Crystallogr D Struct Biol 80: 605-619
- PubMed: 39012716 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798324006557
- Primary Citation Related Structures:
8OPU, 8OPV, 8OPW, 8OPX, 8OPY, 8OQL, 8OQM, 8OQN, 8OQO, 8OQP, 8OQQ, 8OQR, 8OQS, 8OQT, 8OQU, 8OQV, 8PF8 - PubMed Abstract:
The Mycobacterium tuberculosis trifunctional enzyme (MtTFE) is an α 2 β 2 tetrameric enzyme in which the α-chain harbors the 2E-enoyl-CoA hydratase (ECH) and 3S-hydroxyacyl-CoA dehydrogenase (HAD) active sites, and the β-chain provides the 3-ketoacyl-CoA thiolase (KAT) active site. Linear, medium-chain and long-chain 2E-enoyl-CoA molecules are the preferred substrates of MtTFE. Previous crystallographic binding and modeling studies identified binding sites for the acyl-CoA substrates at the three active sites, as well as the NAD binding pocket at the HAD active site. These studies also identified three additional CoA binding sites on the surface of MtTFE that are different from the active sites. It has been proposed that one of these additional sites could be of functional relevance for the substrate channeling (by surface crawling) of reaction intermediates between the three active sites. Here, 226 fragments were screened in a crystallographic fragment-binding study of MtTFE crystals, resulting in the structures of 16 MtTFE-fragment complexes. Analysis of the 121 fragment-binding events shows that the ECH active site is the `binding hotspot' for the tested fragments, with 41 binding events. The mode of binding of the fragments bound at the active sites provides additional insight into how the long-chain acyl moiety of the substrates can be accommodated at their proposed binding pockets. In addition, the 20 fragment-binding events between the active sites identify potential transient binding sites of reaction intermediates relevant to the possible channeling of substrates between these active sites. These results provide a basis for further studies to understand the functional relevance of the latter binding sites and to identify substrates for which channeling is crucial.
- Faculty of Biochemistry and Molecular Medicine, University of Oulu, Oulu, Finland.
Organizational Affiliation: