Structure-Guided Design and Synthesis of a Pyridazinone Series of Trypanosoma cruzi Proteasome Inhibitors.
Thomas, M.G., McGonagle, K., Rowland, P., Robinson, D.A., Dodd, P.G., Camino-Diaz, I., Campbell, L., Cantizani, J., Castaneda, P., Conn, D., Craggs, P.D., Edwards, D., Ferguson, L., Fosberry, A., Frame, L., Goswami, P., Hu, X., Korczynska, J., MacLean, L., Martin, J., Mutter, N., Osuna-Cabello, M., Paterson, C., Pena, I., Pinto, E.G., Pont, C., Riley, J., Shishikura, Y., Simeons, F.R.C., Stojanovski, L., Thomas, J., Wrobel, K., Young, R.J., Zmuda, F., Zuccotto, F., Read, K.D., Gilbert, I.H., Marco, M., Miles, T.J., Manzano, P., De Rycker, M.(2023) J Med Chem 66: 10413-10431
- PubMed: 37506194 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.3c00582
- Primary Citation Related Structures:
8OLU - PubMed Abstract:
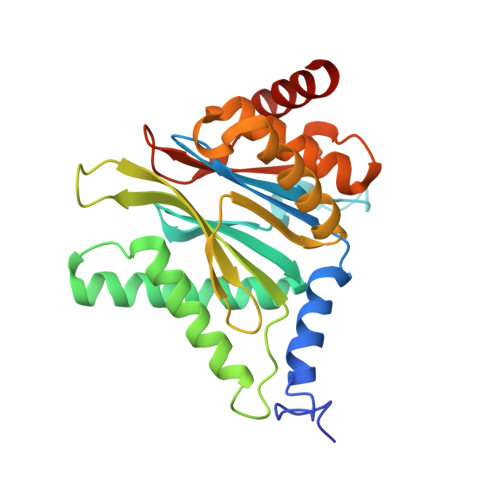
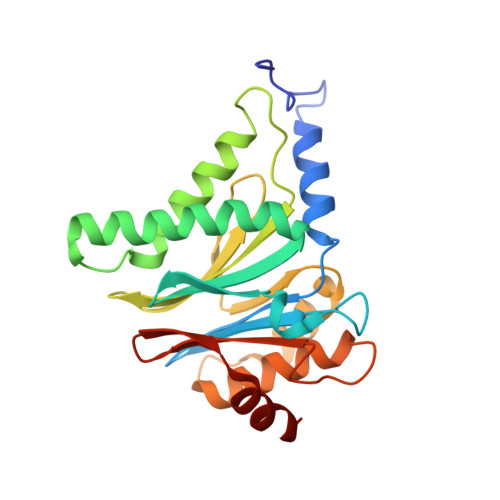
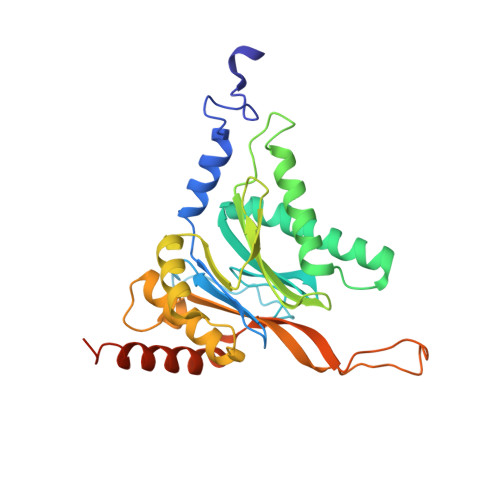
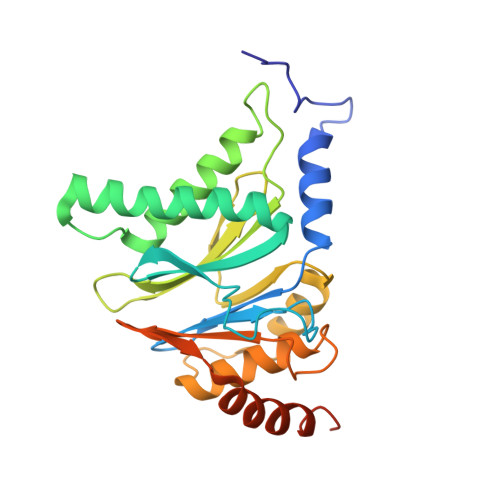
There is an urgent need for new treatments for Chagas disease, a parasitic infection which mostly impacts South and Central America. We previously reported on the discovery of GSK3494245/DDD01305143, a preclinical candidate for visceral leishmaniasis which acted through inhibition of the Leishmania proteasome. A related analogue, active against Trypanosoma cruzi , showed suboptimal efficacy in an animal model of Chagas disease, so alternative proteasome inhibitors were investigated. Screening a library of phenotypically active analogues against the T. cruzi proteasome identified an active, selective pyridazinone, the development of which is described herein. We obtained a cryo-EM co-structure of proteasome and a key inhibitor and used this to drive optimization of the compounds. Alongside this, optimization of the absorption, distribution, metabolism, and excretion (ADME) properties afforded a suitable compound for mouse efficacy studies. The outcome of these studies is discussed, alongside future plans to further understand the series and its potential to deliver a new treatment for Chagas disease.
- Drug Discovery Unit, University of Dundee, School of Life Sciences, Dow Street, Dundee, U.K., DD1 5EH.
Organizational Affiliation: