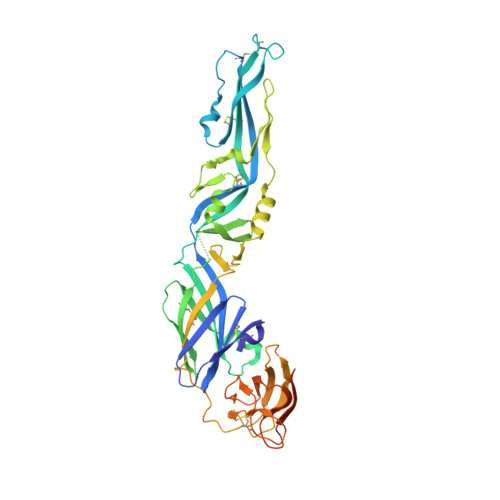
New insight into flavivirus maturation from structure/function studies of the yellow fever virus envelope protein complex.
Crampon, E., Covernton, E., Vaney, M.C., Dellarole, M., Sommer, S., Sharma, A., Haouz, A., England, P., Lepault, J., Duquerroy, S., Rey, F.A., Barba-Spaeth, G.(2023) mBio 14: e0070623-e0070623
- PubMed: 37607061 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mbio.00706-23
- Primary Citation Related Structures:
8OFN - PubMed Abstract:
All enveloped viruses enter cells by fusing their envelope with a target cell membrane while avoiding premature fusion with membranes of the producer cell-the latter being particularly important for viruses that bud at internal membranes. Flaviviruses bud in the endoplasmic reticulum, are transported through the TGN to reach the external milieu, and enter other cells via receptor-mediated endocytosis. The trigger for membrane fusion is the acidic environment of early endosomes, which has a similar pH to the TGN of the producer cell. The viral particles therefore become activated to react to mildly acidic pH only after their release into the neutral pH extracellular environment. Our study shows that for yellow fever virus (YFV), the mechanism of activation involves actively knocking out the fusion brake (protein pr) through a localized conformational change of the envelope protein upon exposure to the neutral pH external environment. Our study has important implications for understanding the molecular mechanism of flavivirus fusion activation in general and points to an alternative way of interfering with this process as an antiviral treatment.
- Institut Pasteur, Université Paris Cité, CNRS UMR 3569, Unité de Virologie Structurale , Paris, France.
Organizational Affiliation: