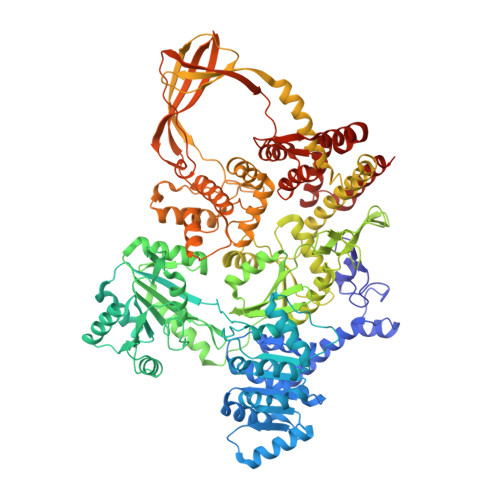
Structure of reverse gyrase with a minimal latch that supports ATP-dependent positive supercoiling without specific interactions with the topoisomerase domain.
Mhaindarkar, V.P., Rasche, R., Kummel, D., Rudolph, M.G., Klostermeier, D.(2023) Acta Crystallogr D Struct Biol 79: 498-507
- PubMed: 37204816 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798323002565
- Primary Citation Related Structures:
7FSE, 7FSF, 8OFB - PubMed Abstract:
Reverse gyrase is the only topoisomerase that introduces positive supercoils into DNA in an ATP-dependent reaction. Positive DNA supercoiling becomes possible through the functional cooperation of the N-terminal helicase domain of reverse gyrase with its C-terminal type IA topoisomerase domain. This cooperation is mediated by a reverse-gyrase-specific insertion into the helicase domain termed the `latch'. The latch consists of a globular domain inserted at the top of a β-bulge loop that connects this globular part to the helicase domain. While the globular domain shows little conservation in sequence and length and is dispensable for DNA supercoiling, the β-bulge loop is required for supercoiling activity. It has previously been shown that the β-bulge loop constitutes a minimal latch that couples ATP-dependent processes in the helicase domain to DNA processing by the topoisomerase domain. Here, the crystal structure of Thermotoga maritima reverse gyrase with such a β-bulge loop as a minimal latch is reported. It is shown that the β-bulge loop supports ATP-dependent DNA supercoiling of reverse gyrase without engaging in specific interactions with the topoisomerase domain. When only a small latch or no latch is present, a helix in the nearby helicase domain of T. maritima reverse gyrase partially unfolds. Comparison of the sequences and predicted structures of latch regions in other reverse gyrases shows that neither sequence nor structure are decisive factors for latch functionality; instead, the decisive factors are likely to be electrostatics and plain steric bulk.
- Institute for Physical Chemistry, University of Muenster, Corrensstrasse 30, 48149 Muenster, Germany.
Organizational Affiliation: