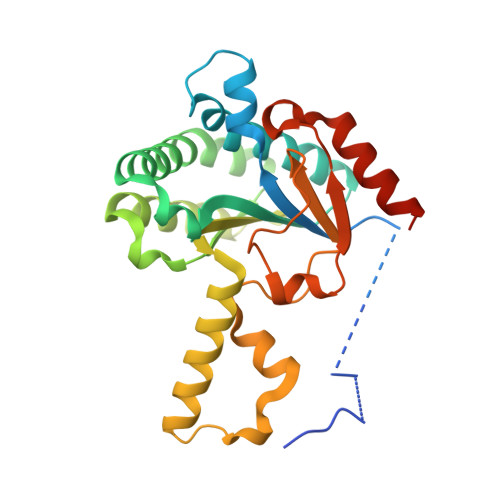
Biosynthesising various rubber-like polymers by reconstituting prenyltransferases on Hevea rubber particles: molecular and structural bases of the versatile system
Suenaga-Hiromori, M., Ishii, T., Imaizumi, R., Takeshita, K., Yanai, T., Matsuura, H., Sakai, N., Mikami, T., Takahashi, T., Minakawa, C., Yamaguchi, H., Yanbe, F., Waki, T., Tozawa, Y., Miyagi-Inoue, Y., Fushihara, K., Yamamoto, M., Kataoka, K., Nakayama, T., Yamashita, S., Takahashi, S.To be published.