Crystal Structure of p97-N/D1 Hexamer Complexed with FAF1 UBX Domain
Kang, W.(2024) J Korean Chem Soc
Experimental Data Snapshot
Starting Model: experimental
View more details
(2024) J Korean Chem Soc
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Transitional endoplasmic reticulum ATPase | 438 | Homo sapiens | Mutation(s): 3 Gene Names: VCP EC: 3.6.4.6 | 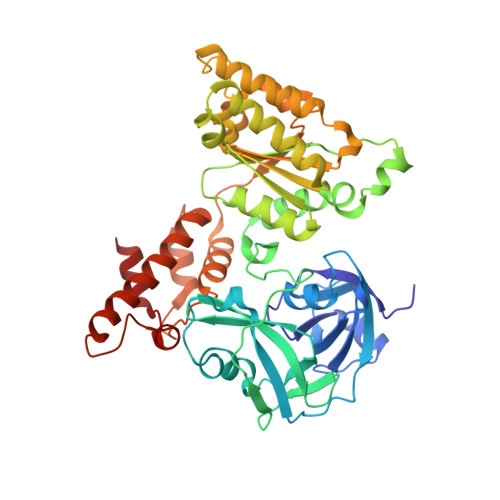 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P55072 GTEx: ENSG00000165280 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P55072 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| FAS-associated factor 1 | 76 | Homo sapiens | Mutation(s): 0 Gene Names: FAF1 | 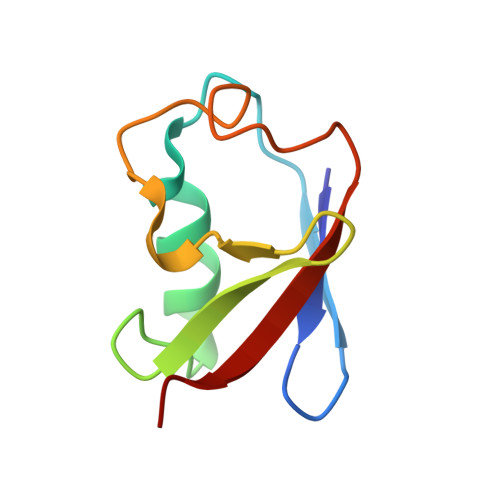 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9UNN5 GTEx: ENSG00000185104 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9UNN5 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ADP (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | AA [auth C] BA [auth D] CA [auth E] DA [auth F] EA [auth G] | ADENOSINE-5'-DIPHOSPHATE C10 H15 N5 O10 P2 XTWYTFMLZFPYCI-KQYNXXCUSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 106.665 | α = 71.52 |
| b = 134.484 | β = 80.87 |
| c = 148.223 | γ = 87.53 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Research Foundation (NRF, Korea) | Korea, Republic Of | 2021R1A6A1A10044154 |
| National Research Foundation (NRF, Korea) | Korea, Republic Of | 2022R1C1C1004221 |