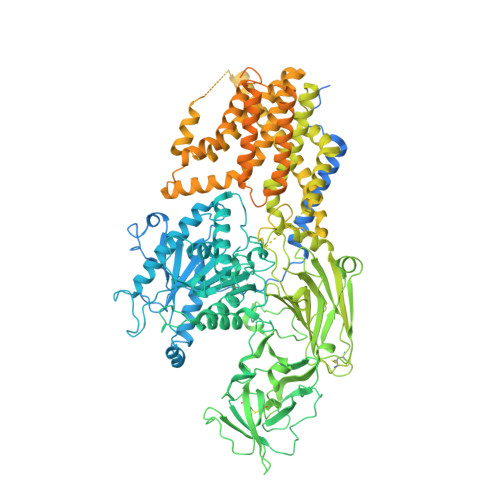
Molecular basis of the inositol deacylase PGAP1 involved in quality control of GPI-AP biogenesis.
Hong, J., Li, T., Chao, Y., Xu, Y., Zhu, Z., Zhou, Z., Gu, W., Qu, Q., Li, D.(2024) Nat Commun 15: 8-8
- PubMed: 38167496 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-44568-2
- Primary Citation Related Structures:
8K9Q, 8K9R, 8K9T - PubMed Abstract:
The secretion and quality control of glycosylphosphatidylinositol-anchored proteins (GPI-APs) necessitates post-attachment remodeling initiated by the evolutionarily conserved PGAP1, which deacylates the inositol in nascent GPI-APs. Impairment of PGAP1 activity leads to developmental diseases in humans and fatality and infertility in animals. Here, we present three PGAP1 structures (2.66-2.84 Å), revealing its 10-transmembrane architecture and product-enzyme interaction details. PGAP1 holds GPI-AP acyl chains in an optimally organized, guitar-shaped cavity with apparent energetic penalties from hydrophobic-hydrophilic mismatches. However, abundant glycan-mediated interactions in the lumen counterbalance these repulsions, likely conferring substrate fidelity and preventing off-target hydrolysis of bulk membrane lipids. Structural and biochemical analyses uncover a serine hydrolase-type catalysis with atypical features and imply mechanisms for substrate entrance and product release involving a drawing compass movement of GPI-APs. Our findings advance the mechanistic understanding of GPI-AP remodeling.
- State Key Laboratory of Molecular Biology, Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences; University of Chinese Academy of Sciences, 320 Yueyang Road, Shanghai, 200031, China.
Organizational Affiliation: