Potent human neutralizing antibodies against Nipah virus derived from two ancestral antibody heavy chains.
Chen, L., Sun, M., Zhang, H., Zhang, X., Yao, Y., Li, M., Li, K., Fan, P., Zhang, H., Qin, Y., Zhang, Z., Li, E., Chen, Z., Guan, W., Li, S., Yu, C., Zhang, K., Gong, R., Chiu, S.(2024) Nat Commun 15: 2987-2987
- PubMed: 38582870 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-47213-8
- Primary Citation Related Structures:
8K3C - PubMed Abstract:
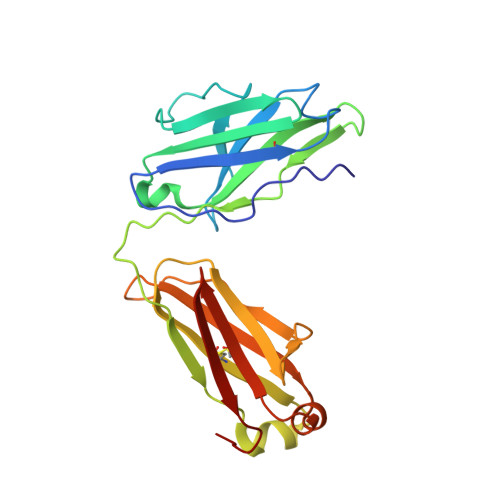
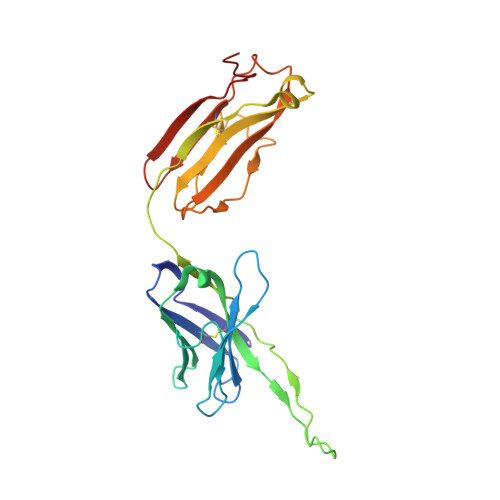
Nipah virus (NiV) is a World Health Organization priority pathogen and there are currently no approved drugs for clinical immunotherapy. Through the use of a naïve human phage-displayed Fab library, two neutralizing antibodies (NiV41 and NiV42) targeting the NiV receptor binding protein (RBP) were identified. Following affinity maturation, antibodies derived from NiV41 display cross-reactivity against both NiV and Hendra virus (HeV), whereas the antibody based on NiV42 is only specific to NiV. Results of immunogenetic analysis reveal a correlation between the maturation of antibodies and their antiviral activity. In vivo testing of NiV41 and its mature form (41-6) show protective efficacy against a lethal NiV challenge in hamsters. Furthermore, a 2.88 Å Cryo-EM structure of the tetrameric RBP and antibody complex demonstrates that 41-6 blocks the receptor binding interface. These findings can be beneficial for the development of antiviral drugs and the design of vaccines with broad spectrum against henipaviruses.
- CAS Key Laboratory of Special Pathogens and Biosafety, Wuhan Institute of Virology, Center for Biosafety Mega-Science, Chinese Academy of Sciences, Wuhan, Hubei, China.
Organizational Affiliation: