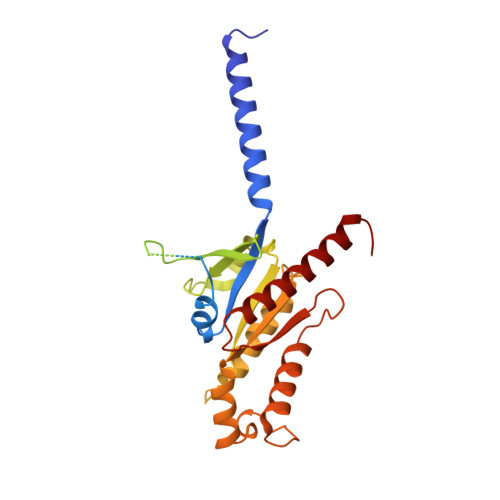
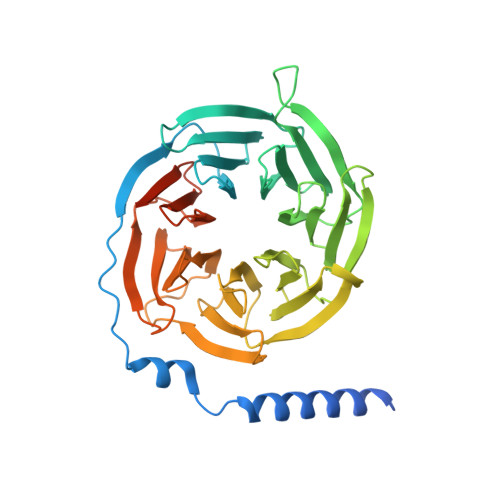
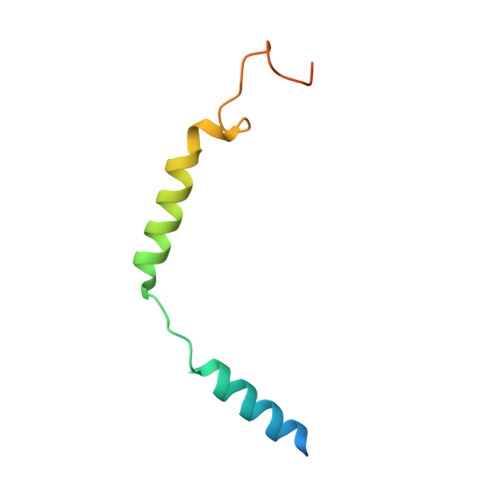
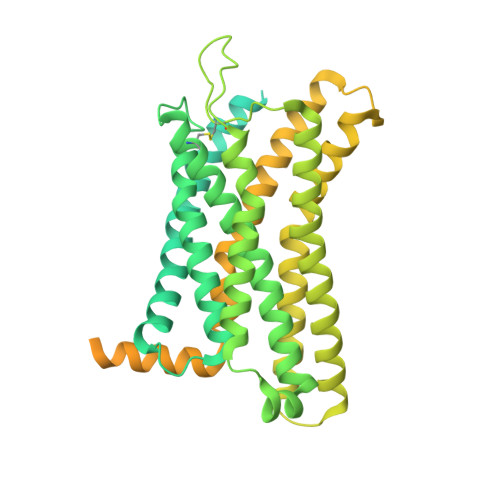
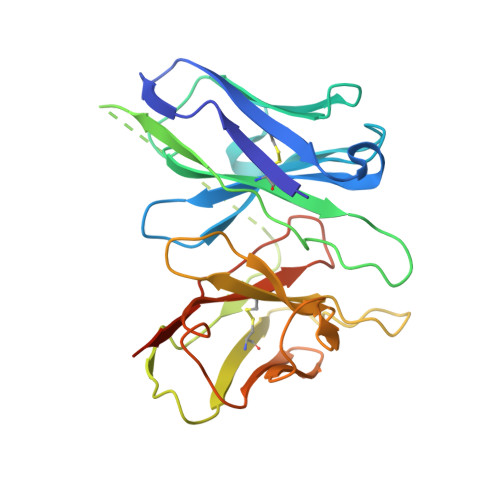
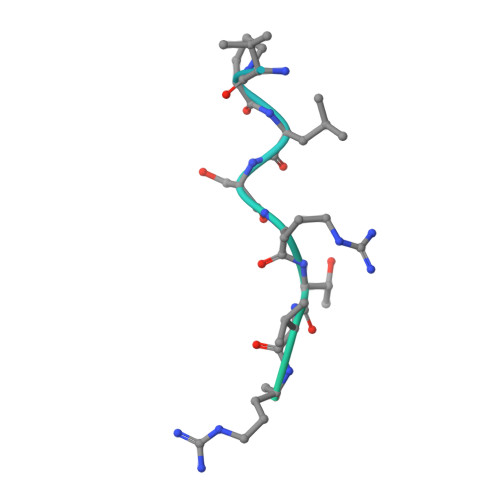
Structure basis for the modulation of CXC chemokine receptor 3 by antagonist AMG487.
Jiao, H., Pang, B., Chiang, Y.C., Chen, Q., Pan, Q., Ren, R., Hu, H.(2023) Cell Discov 9: 119-119
- PubMed: 38012179 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-023-00617-0
- Primary Citation Related Structures:
8K2W, 8K2X - Kobilka Institute of Innovative Drug Discovery, School of Medicine, The Chinese University of Hong Kong, Shenzhen, Shenzhen, Guangdong, China.
Organizational Affiliation: