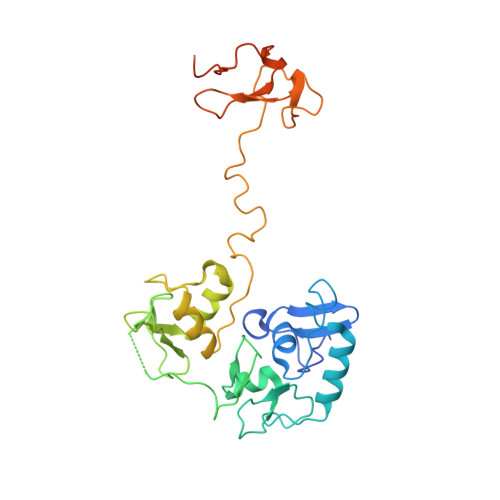
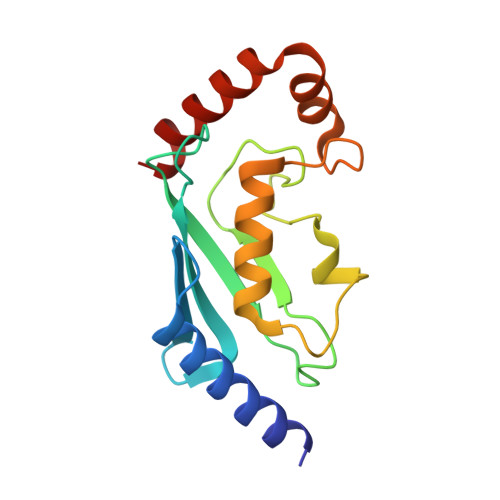
Molecular basis for PHF7-mediated ubiquitination of histone H3.
Lee, H.S., Bang, I., You, J., Jeong, T.K., Kim, C.R., Hwang, M., Kim, J.S., Baek, S.H., Song, J.J., Choi, H.J.(2023) Genes Dev 37: 984-997
- PubMed: 37993255 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/gad.350989.123
- Primary Citation Related Structures:
8JWJ, 8JWS, 8JWU - PubMed Abstract:
The RING-type E3 ligase has been known for over two decades, yet its diverse modes of action are still the subject of active research. Plant homeodomain (PHD) finger protein 7 (PHF7) is a RING-type E3 ubiquitin ligase responsible for histone ubiquitination. PHF7 comprises three zinc finger domains: an extended PHD (ePHD), a RING domain, and a PHD. While the function of the RING domain is largely understood, the roles of the other two domains in E3 ligase activity remain elusive. Here, we present the crystal structure of PHF7 in complex with the E2 ubiquitin-conjugating enzyme (E2). Our structure shows that E2 is effectively captured between the RING domain and the C-terminal PHD, facilitating E2 recruitment through direct contact. In addition, through in vitro binding and functional assays, we demonstrate that the N-terminal ePHD recognizes the nucleosome via DNA binding, whereas the C-terminal PHD is involved in histone H3 recognition. Our results provide a molecular basis for the E3 ligase activity of PHF7 and uncover the specific yet collaborative contributions of each domain to the PHF7 ubiquitination activity.
- Department of Biological Sciences, Seoul National University, Seoul 08826, Republic of Korea.
Organizational Affiliation: