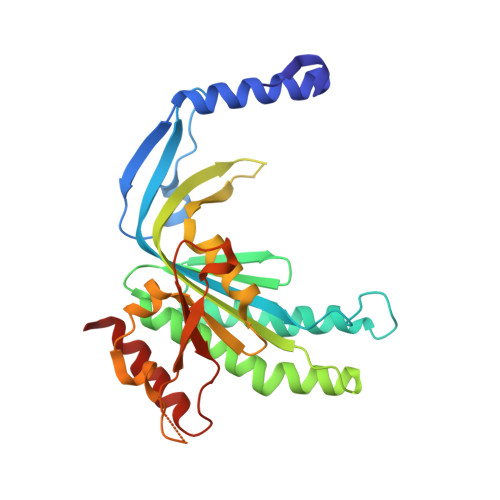
Structural and functional characterization of cyclic pyrimidine-regulated anti-phage system.
Hou, M.H., Chen, C.J., Yang, C.S., Wang, Y.C., Chen, Y.(2024) Nat Commun 15: 5634-5634
- PubMed: 38965224 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-49861-2
- Primary Citation Related Structures:
8JSF, 8JSJ, 8JSK, 8JSZ - PubMed Abstract:
3',5'-cyclic uridine monophosphate (cUMP) and 3',5'-cyclic cytidine monophosphate (cCMP) have been established as bacterial second messengers in the phage defense system, named pyrimidine cyclase system for anti-phage resistance (Pycsar). This system consists of a pyrimidine cyclase and a cyclic pyrimidine receptor protein. However, the molecular mechanism underlying cyclic pyrimidine synthesis and recognition remains unclear. Herein, we determine the crystal structures of a uridylate cyclase and a cytidylate cyclase, revealing the conserved residues for cUMP and cCMP production, respectively. In addition, a distinct zinc-finger motif of the uridylate cyclase is identified to confer substantial resistance against phage infections. Furthermore, structural characterization of cUMP receptor protein PycTIR provides clear picture of specific cUMP recognition and identifies a conserved N-terminal extension that mediates PycTIR oligomerization and activation. Overall, our results contribute to the understanding of cyclic pyrimidine-mediated bacterial defense.
- Department of Food Science and Biotechnology, National Chung Hsing University, Taichung, 40227, Taiwan.
Organizational Affiliation: