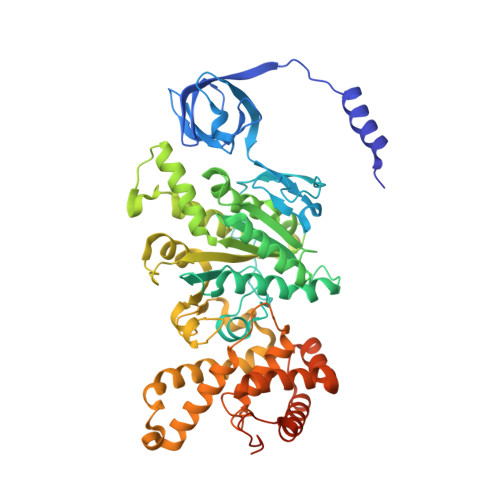
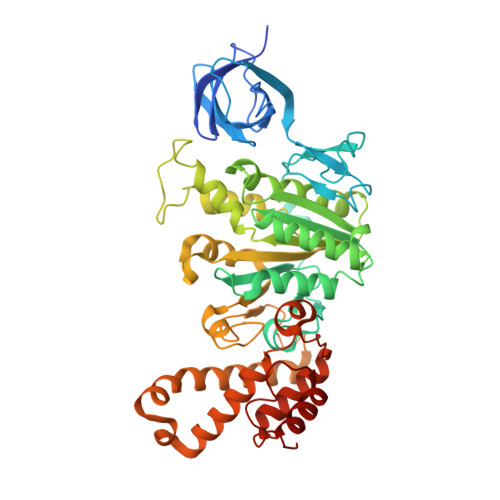
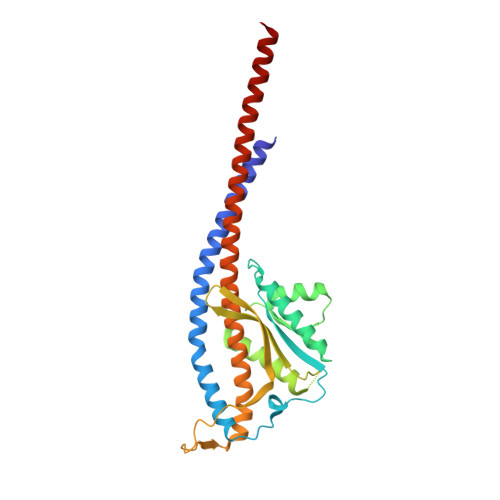
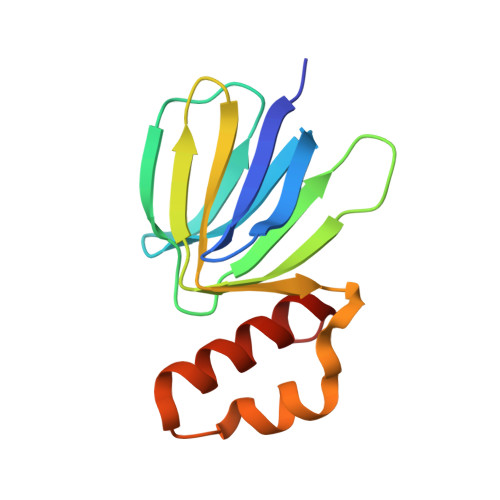
Inhibition of M. tuberculosis and human ATP synthase by BDQ and TBAJ-587.
Zhang, Y., Lai, Y., Zhou, S., Ran, T., Zhang, Y., Zhao, Z., Feng, Z., Yu, L., Xu, J., Shi, K., Wang, J., Pang, Y., Li, L., Chen, H., Guddat, L.W., Gao, Y., Liu, F., Rao, Z., Gong, H.(2024) Nature 631: 409-414
- PubMed: 38961288 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-024-07605-8
- Primary Citation Related Structures:
8J0S, 8J0T, 8J57, 8J58, 8JR0, 8JR1, 8KHF, 8KI3 - PubMed Abstract:
Bedaquiline (BDQ), a first-in-class diarylquinoline anti-tuberculosis drug, and its analogue, TBAJ-587, prevent the growth and proliferation of Mycobacterium tuberculosis by inhibiting ATP synthase 1,2 . However, BDQ also inhibits human ATP synthase 3 . At present, how these compounds interact with either M. tuberculosis ATP synthase or human ATP synthase is unclear. Here we present cryogenic electron microscopy structures of M. tuberculosis ATP synthase with and without BDQ and TBAJ-587 bound, and human ATP synthase bound to BDQ. The two inhibitors interact with subunit a and the c-ring at the leading site, c-only sites and lagging site in M. tuberculosis ATP synthase, showing that BDQ and TBAJ-587 have similar modes of action. The quinolinyl and dimethylamino units of the compounds make extensive contacts with the protein. The structure of human ATP synthase in complex with BDQ reveals that the BDQ-binding site is similar to that observed for the leading site in M. tuberculosis ATP synthase, and that the quinolinyl unit also interacts extensively with the human enzyme. This study will improve researchers' understanding of the similarities and differences between human ATP synthase and M. tuberculosis ATP synthase in terms of the mode of BDQ binding, and will allow the rational design of novel diarylquinolines as anti-tuberculosis drugs.
- State Key Laboratory of Medicinal Chemical Biology and College of Life Sciences, Nankai University, Tianjin, China.
Organizational Affiliation: