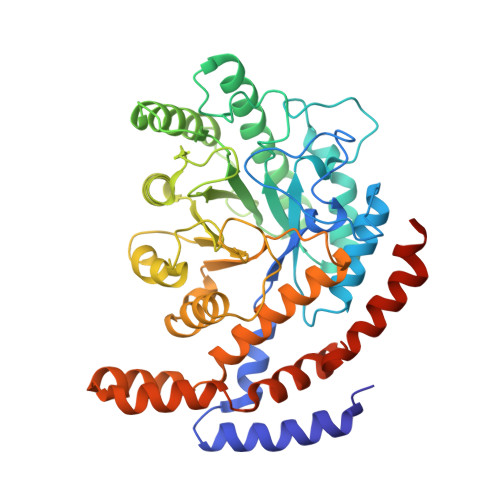
X-ray structure and characterization of a probiotic Lactobacillus rhamnosus Probio-M9 L-rhamnose isomerase.
Yoshida, H., Yamamoto, N., Kurahara, L.H., Izumori, K., Yoshihara, A.(2024) Appl Microbiol Biotechnol 108: 249-249
- PubMed: 38430263 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s00253-024-13075-9
- Primary Citation Related Structures:
8JQ3, 8JQ4, 8JQ5, 8JQ6 - PubMed Abstract:
A recombinant L-rhamnose isomerase (L-RhI) from probiotic Lactobacillus rhamnosus Probio-M9 (L. rhamnosus Probio-M9) was expressed. L. rhamnosus Probio-M9 was isolated from human colostrum and identified as a probiotic lactic acid bacterium, which can grow using L-rhamnose. L-RhI is one of the enzymes involved in L-rhamnose metabolism and catalyzes the reversible isomerization between L-rhamnose and L-rhamnulose. Some L-RhIs were reported to catalyze isomerization not only between L-rhamnose and L-rhamnulose but also between D-allulose and D-allose, which are known as rare sugars. Those L-RhIs are attractive enzymes for rare sugar production and have the potential to be further improved by enzyme engineering; however, the known crystal structures of L-RhIs recognizing rare sugars are limited. In addition, the optimum pH levels of most reported L-RhIs are basic rather than neutral, and such a basic condition causes non-enzymatic aldose-ketose isomerization, resulting in unexpected by-products. Herein, we report the crystal structures of L. rhamnosus Probio-M9 L-RhI (LrL-RhI) in complexes with L-rhamnose, D-allulose, and D-allose, which show enzyme activity toward L-rhamnose, D-allulose, and D-allose in acidic conditions, though the activity toward D-allose was low. In the complex with L-rhamnose, L-rhamnopyranose was found in the catalytic site, showing favorable recognition for catalysis. In the complex with D-allulose, D-allulofuranose and ring-opened D-allulose were observed in the catalytic site. However, bound D-allose in the pyranose form was found in the catalytic site of the complex with D-allose, which was unfavorable for recognition, like an inhibition mode. The structure of the complex may explain the low activity toward D-allose. KEY POINTS: • Crystal structures of LrL-RhI in complexes with substrates were determined. • LrL-RhI exhibits enzyme activity toward L-rhamnose, D-allulose, and D-allose. • The LrL-RhI is active in acidic conditions.
- Department of Basic Life Science, Faculty of Medicine, Kagawa University, 1750-1 Ikenobe, Miki-Cho, Kita-Gun, Kagawa, 761-0793, Japan. yoshida.hiromi@kagawa-u.ac.jp.
Organizational Affiliation: