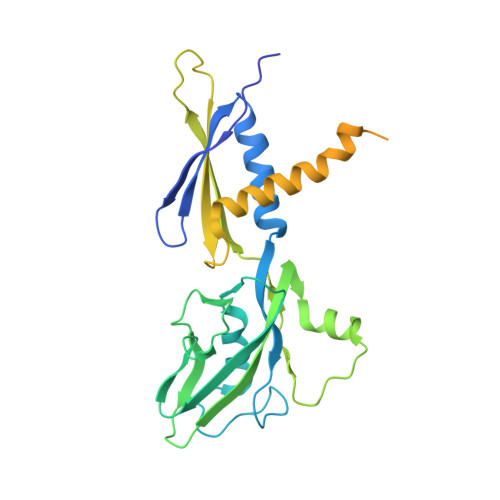
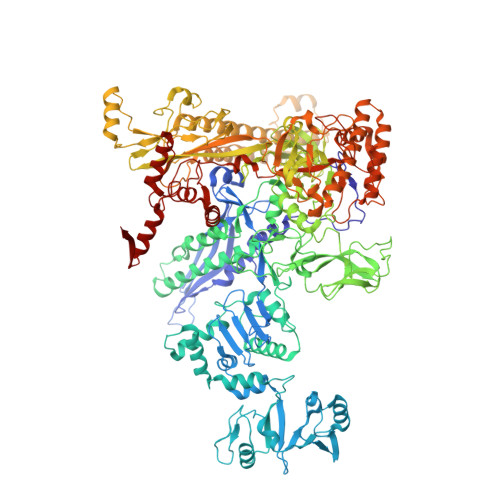
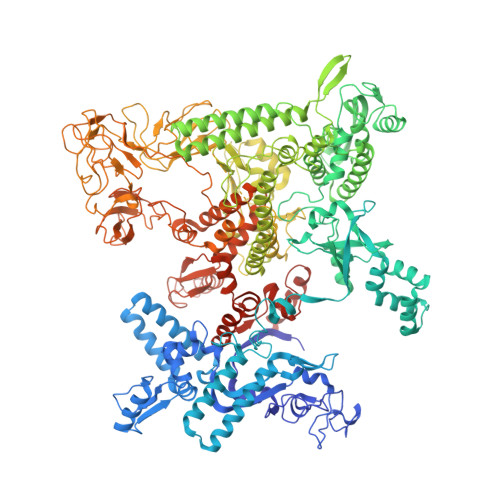
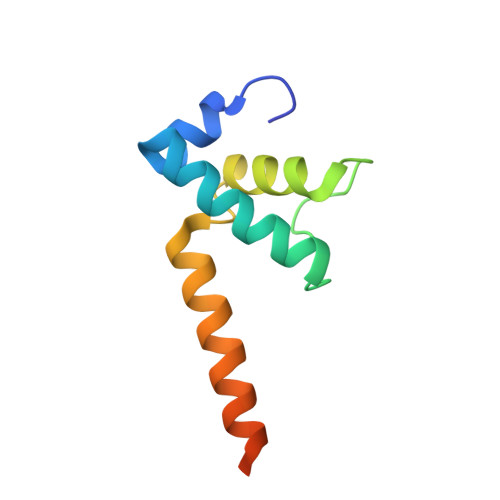
Structural basis of transcriptional activation by the OmpR/PhoB-family response regulator PmrA.
Lou, Y.C., Huang, H.Y., Yeh, H.H., Chiang, W.H., Chen, C., Wu, K.P.(2023) Nucleic Acids Res 51: 10049-10058
- PubMed: 37665001 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkad724
- Primary Citation Related Structures:
8JO2 - PubMed Abstract:
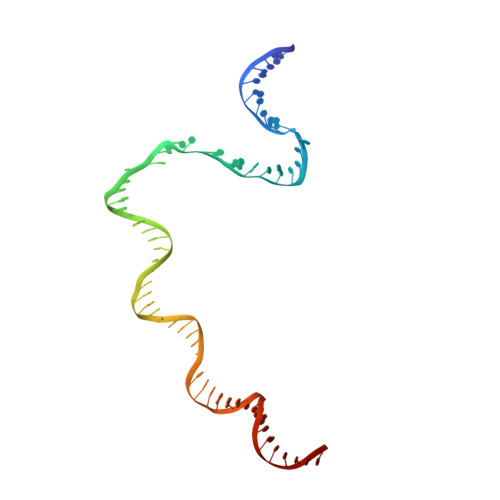
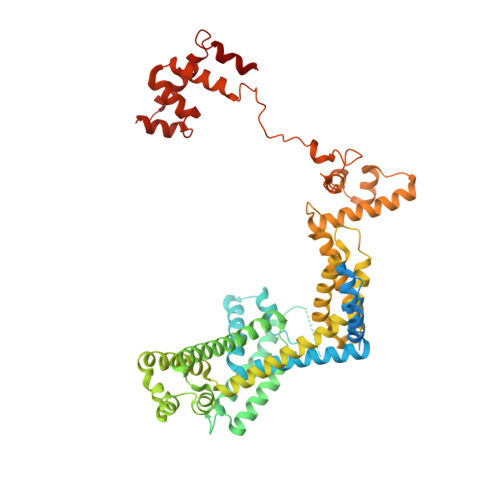
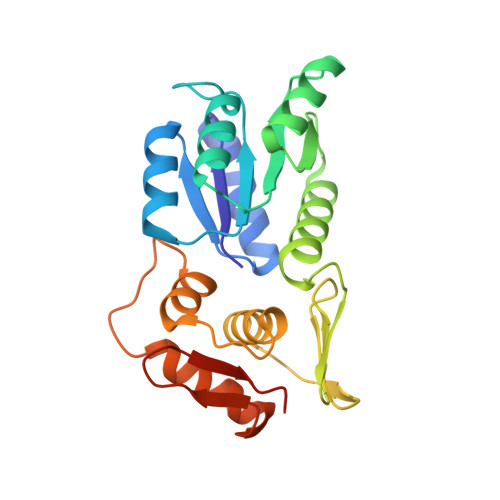
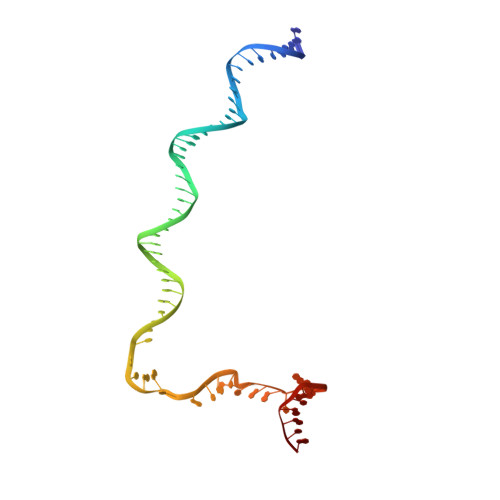
PmrA, an OmpR/PhoB-family response regulator, triggers gene transcription responsible for polymyxin resistance in bacteria by recognizing promoters where the canonical-35 element is replaced by the pmra-box, representing the PmrA recognition sequence. Here, we report a cryo-electron microscopy (cryo-EM) structure of a bacterial PmrA-dependent transcription activation complex (TAC) containing a PmrA dimer, an RNA polymerase σ70 holoenzyme (RNAPH) and the pbgP promoter DNA. Our structure reveals that the RNAPH mainly contacts the PmrA C-terminal DNA-binding domain (DBD) via electrostatic interactions and reorients the DBD three base pairs upstream of the pmra-box, resulting in a dynamic TAC conformation. In vivo assays show that the substitution of the DNA-recognition residue eliminated its transcriptional activity, while variants with altered RNAPH-interacting residues resulted in enhanced transcriptional activity. Our findings suggest that both PmrA recognition-induced DNA distortion and PmrA promoter escape play crucial roles in its transcriptional activation.
- Biomedical Translation Research Center, Academia Sinica, Taipei 11529, Taiwan.
Organizational Affiliation: