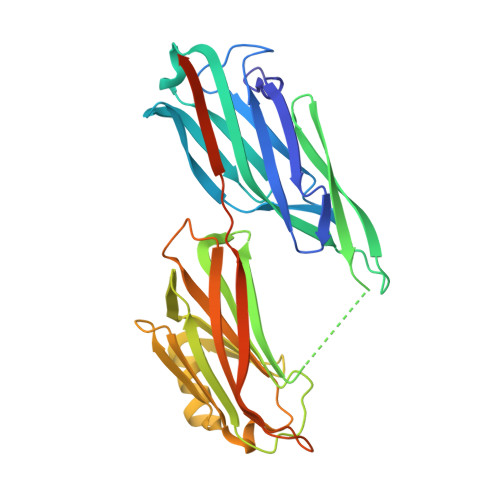
Structure, Stability and Binding Properties of Collagen-Binding Domains from Streptococcus mutans.
Nishi, A., Matsui, H., Hirata, A., Mukaiyama, A., Tanaka, S.-i., Yoshizawa, T., Matsumura, H., Nomura, R., Nakano, K., Takano, K.(2023) Chemistry 5: 1911-1920