Structure-based design and synthesis of BML284 derivatives: A novel class of colchicine-site noncovalent tubulin degradation agents.
Zhang, C., Yan, W., Liu, Y., Tang, M., Teng, Y., Wang, F., Hu, X., Zhao, M., Yang, J., Li, Y.(2024) Eur J Med Chem 268: 116265-116265
- PubMed: 38430854 Search on PubMed
- DOI: https://doi.org/10.1016/j.ejmech.2024.116265
- Primary Citation Related Structures:
8JJB, 8JJC - PubMed Abstract:
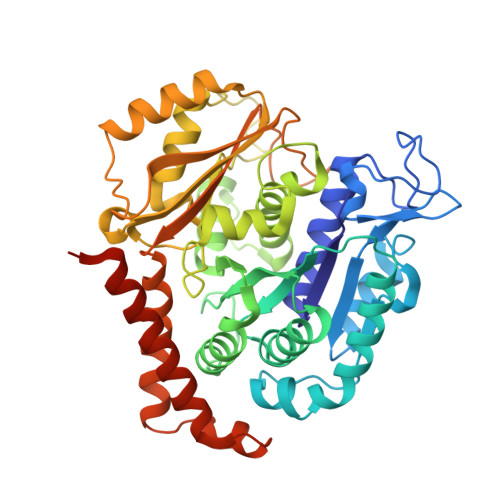
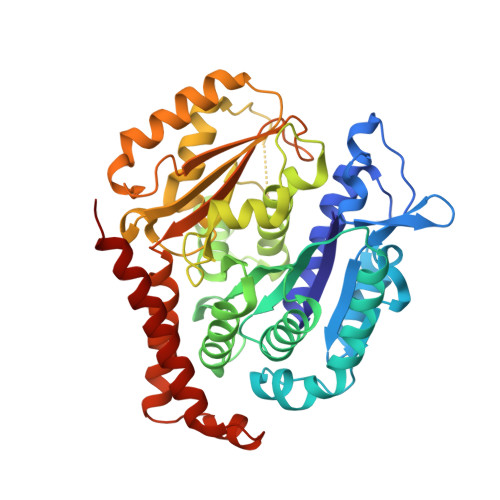
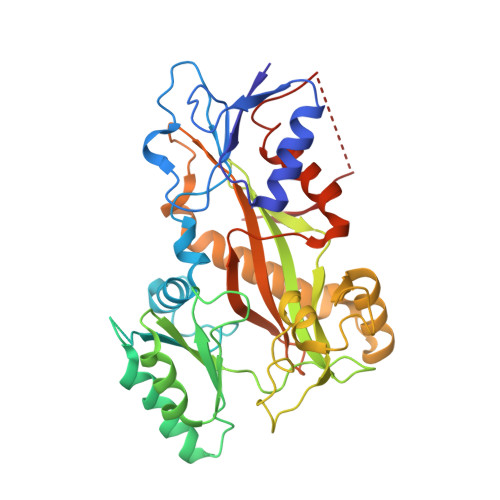
Our previous studies have demonstrated that BML284 is a colchicine-site tubulin degradation agent. To improve its antiproliferative properties, 45 derivatives or analogs of BML284 were designed and synthesized based on the cocrystal structure of BML284 and tubulin. Among them, 5i was the most potent derivative, with IC 50 values ranging from 0.02 to 0.05 μM against the five tested tumor cell lines. Structure-activity relationship studies verified that the N1 atom of the pyrimidine ring was the key functional group for its tubulin degradation ability. The 5i-tubulin cocrystal complex revealed that the binding pattern of 5i to tubulin is similar to that of BML284. However, replacing the benzodioxole ring with an indole ring strengthened the hydrogen bond formed by the 2-amino group with E198, which improved the antiproliferative activity of 5i. Compound 5i effectively suppressed tumor growth at an intravenous dose of 40 mg/kg (every 2 days) in paclitaxel sensitive A2780S and paclitaxel resistant A2780T ovarian xenograft models, with tumor growth inhibition values of 79.4% and 82.0%, respectively, without apparent side effects, showing its potential to overcome multidrug resistance. This study provided a successful example of crystal structure-guided discovery of 5i as a colchicine-targeted tubulin degradation agent, expanding the scope of targeted protein degradation.
- Innovation Center of Nursing Research, Nursing Key Laboratory of Sichuan Province, State Key Laboratory of Biotherapy and Cancer Center, West China Hospital, and Collaborative Innovation Center of Biotherapy, Sichuan University, Chengdu, 610041, Sichuan, China.
Organizational Affiliation: