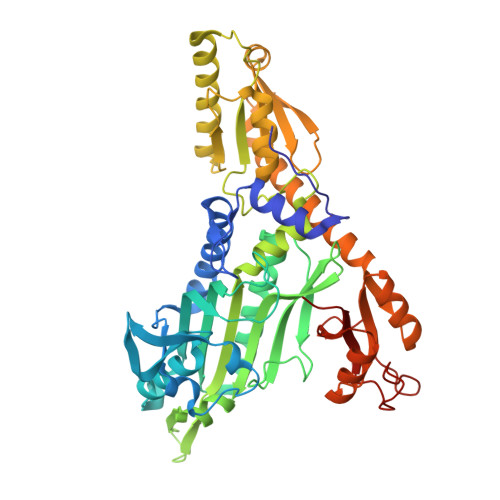
Chemical targeting of prolyl-tRNA synthetase stalls ovarian development and kills malaria vectors.
Goswami, R., Manickam, Y., Goel, J.C., Gupta, S., B M, S., Sharma, A.(2025) J Infect Dis
- PubMed: 40048639 Search on PubMed
- DOI: https://doi.org/10.1093/infdis/jiaf095
- Primary Citation Related Structures:
8JE5, 8JE6, 8JE7 - PubMed Abstract:
Along with rising resistance to antimalarials, the emergence of insecticide resistance in Anopheles mosquito species also remains a serious concern. Here, we reveal two potent compounds that show larvicidal and endectocidal activity against malaria vectors, Anopheles culicifacies and Anopheles stephensi, respectively. We investigated larvicidal activity of two inhibitors against III-instar larvae of Anopheles culicifacies. The survival and fertility of adult female Anopheles stephensi mosquitoes were assessed. Additionally, we purified recombinant prolyl-tRNA synthetase of Anopheles culicifacies and performed enzyme-based assays and structural analysis with the two inhibitors. Our study reveals that the Anopheles culicifacies prolyl-tRNA synthetase (AcProRS) is potently inhibited by halofuginone (HFG) and an ATP mimetic (L95). The evaluation of larvicidal activity of HFG against Anopheles culicifacies III-instar larvae showed a dose-dependent increase in mortality. In adult female Anopheles stephensi mosquitoes, ingestion of HFG via artificial blood feeding resulted in impaired ovary development, reduced egg laying, and decreased overall survival. The potent enzymatic inhibition of AcProRS thus drives the killing of larvae. The co-crystal structure of AcProRS with inhibitors provides a structural basis for improving their potency as future larvicides. Our data suggest the potential for repositioning halofuginone (HFG) and pyrrolidine-based ATP-mimetics (L95) as larvicides. Targeting the vector-encoded aminoacyl-tRNA synthetases provides a new focus for developing effective agents that can control multiple mosquitoe-borne infectious diseases like malaria and dengue.
- ICMR-National Institute of Malaria Research (NIMR); Delhi, India.
Organizational Affiliation: