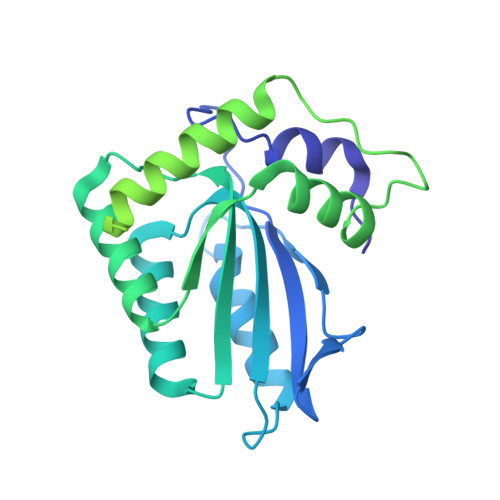
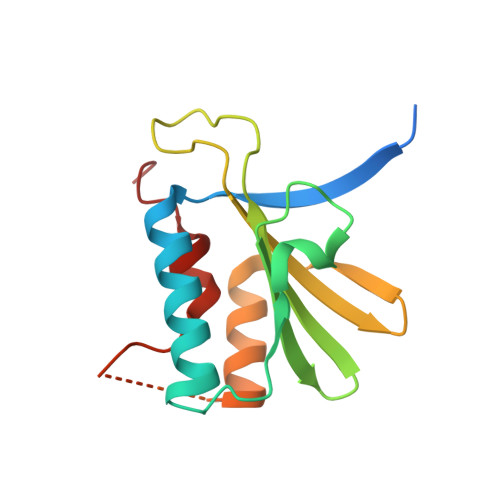
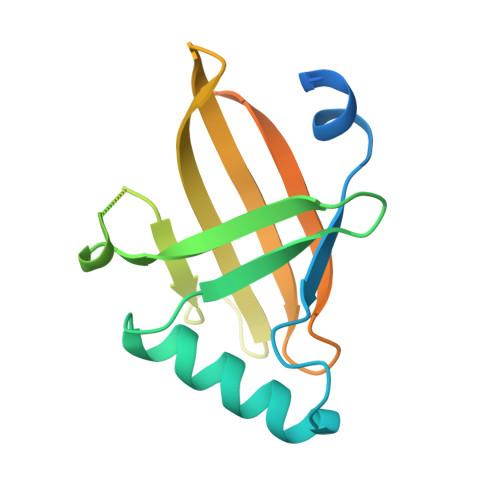
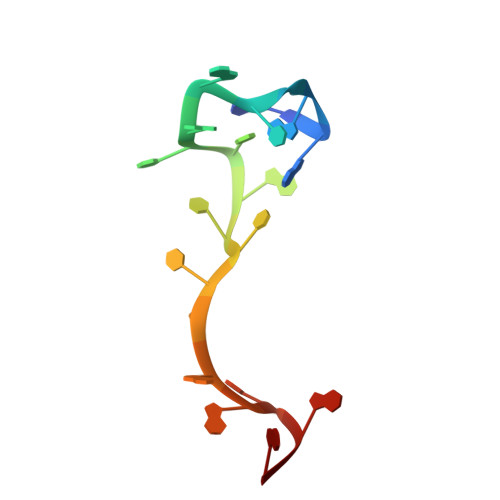
Structural insights into RNA bridging between HIV-1 Vif and antiviral factor APOBEC3G.
Kouno, T., Shibata, S., Shigematsu, M., Hyun, J., Kim, T.G., Matsuo, H., Wolf, M.(2023) Nat Commun 14: 4037-4037
- PubMed: 37419875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-39796-5
- Primary Citation Related Structures:
8H0I, 8J62 - PubMed Abstract:
Great effort has been devoted to discovering the basis of A3G-Vif interaction, the key event of HIV's counteraction mechanism to evade antiviral innate immune response. Here we show reconstitution of the A3G-Vif complex and subsequent A3G ubiquitination in vitro and report the cryo-EM structure of the A3G-Vif complex at 2.8 Å resolution using solubility-enhanced variants of A3G and Vif. We present an atomic model of the A3G-Vif interface, which assembles via known amino acid determinants. This assembly is not achieved by protein-protein interaction alone, but also involves RNA. The cryo-EM structure and in vitro ubiquitination assays identify an adenine/guanine base preference for the interaction and a unique Vif-ribose contact. This establishes the biological significance of an RNA ligand. Further assessment of interactions between A3G, Vif, and RNA ligands show that the A3G-Vif assembly and subsequent ubiquitination can be controlled by amino acid mutations at the interface or by polynucleotide modification, suggesting that a specific chemical moiety would be a promising pharmacophore to inhibit the A3G-Vif interaction.
- Molecular Cryo-Electron Microscopy Unit, Okinawa Institute of Science and Technology Graduate University, 1919-1 Tancha, Onna-son, Okinawa, 904-0495, Japan. 1st.soluble.fla3g@gmail.com.
Organizational Affiliation: