Cryo-EM structure ofABCC4
Chen, Y., Wang, L., Hou, W.T., Zhou, C.Z., Chen, Y., Li, Q.(2023) Nat Cardiovasc Res
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| ATP-binding cassette sub-family C member 4 | 1,325 | Homo sapiens | Mutation(s): 0 Gene Names: ABCC4, MOATB, MRP4 EC: 7.6.2 (PDB Primary Data), 7.6.2.2 (PDB Primary Data), 7.6.2.3 (PDB Primary Data) Membrane Entity: Yes | 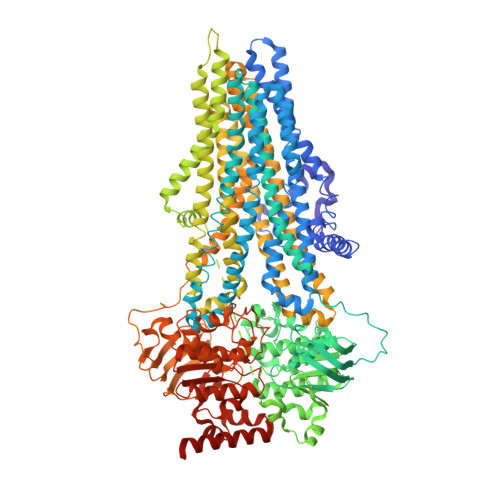 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O15439 GTEx: ENSG00000125257 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O15439 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ATP Download:Ideal Coordinates CCD File | B [auth A], F [auth A] | ADENOSINE-5'-TRIPHOSPHATE C10 H16 N5 O13 P3 ZKHQWZAMYRWXGA-KQYNXXCUSA-N |  | ||
| PUC (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A] | (5Z)-7-{(1R,4S,5S,6R)-6-[(1E,3S)-3-hydroxyoct-1-en-1-yl]-2-oxabicyclo[2.2.1]hept-5-yl}hept-5-enoic acid C21 H34 O4 LQANGKSBLPMBTJ-BRSNVKEHSA-N |  | ||
| MG Download:Ideal Coordinates CCD File | C [auth A], E [auth A] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | 1.18.2_3874: |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Ministry of Science and Technology (MoST, China) | China | 2019YFA0508500 |
| Chinese Academy of Sciences | China | XDB37020202 |