Structural and functional insights into Cdc45 recruitment by Sld7-Sld3 for CMG complex formation
Li, H., Ishizaki, I., Kato, K., Sun, X., Muramatsu, S., Itou, H., Ose, T., Araki, H., Yao, M.(2024) Elife
Experimental Data Snapshot
Starting Models: experimental
View more details
wwPDB Validation 3D Report Full Report
(2024) Elife
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA replication regulator SLD3 | 267 | Saccharomyces cerevisiae S288C | Mutation(s): 0 Gene Names: SLD3 | 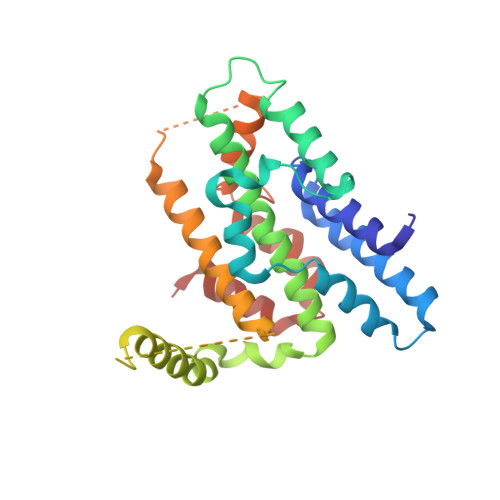 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P53135 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cell division control protein 45 | 650 | Saccharomyces cerevisiae S288C | Mutation(s): 0 Gene Names: CDC45 | 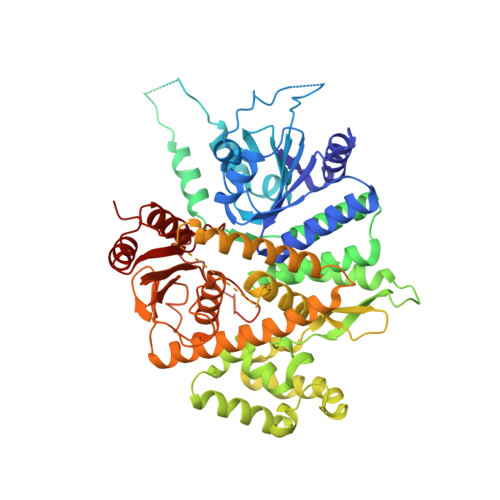 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q08032 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 70.771 | α = 90 |
| b = 107.569 | β = 90 |
| c = 128.283 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XDS | data scaling |
| PHENIX | model building |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Japan Society for the Promotion of Science (JSPS) | Japan | 21H01754 |
| Japan Agency for Medical Research and Development (AMED) | Japan | JP18am0101071 |
| Japan Agency for Medical Research and Development (AMED) | Japan | JP19am0101083 |