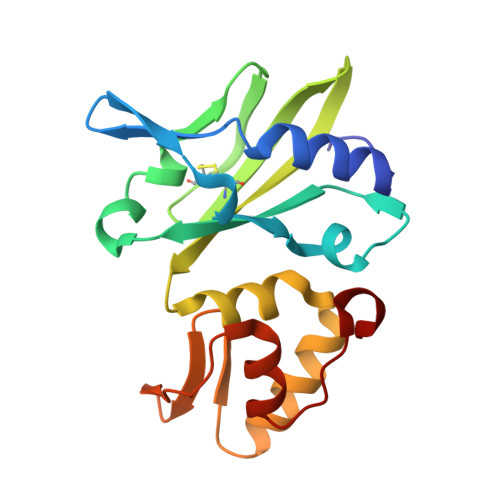
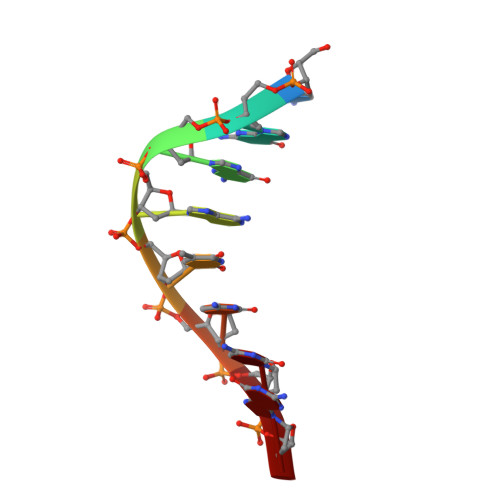
Structural Insight into Polymerase Mechanism via a Chiral Center Generated with a Single Selenium Atom.
Qin, T., Hu, B., Zhao, Q., Wang, Y., Wang, S., Luo, D., Lyu, J., Chen, Y., Gan, J., Huang, Z.(2023) Int J Mol Sci 24
- PubMed: 37958741 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms242115758
- Primary Citation Related Structures:
8ILD, 8ILE, 8ILF, 8ILG, 8ILH, 8ILI - PubMed Abstract:
DNA synthesis catalyzed by DNA polymerase is essential for all life forms, and phosphodiester bond formation with phosphorus center inversion is a key step in this process. Herein, by using a single-selenium-atom-modified dNTP probe, we report a novel strategy to visualize the reaction stereochemistry and catalysis. We capture the before- and after-reaction states and provide explicit evidence of the center inversion and in-line attacking S N 2 mechanism of DNA polymerization, while solving the diastereomer absolute configurations. Further, our kinetic and thermodynamic studies demonstrate that in the presence of Mg 2+ ions (or Mn 2+ ), the binding affinity ( K m ) and reaction selectivity ( k cat / K m ) of dGTPαSe-Rp were 51.1-fold (or 19.5-fold) stronger and 21.8-fold (or 11.3-fold) higher than those of dGTPαSe-Sp, respectively, indicating that the diastereomeric Se-Sp atom was quite disruptive of the binding and catalysis. Our findings reveal that the third metal ion is much more critical than the other two metal ions in both substrate recognition and bond formation, providing insights into how to better design the polymerase inhibitors and discover the therapeutics.
- Key Laboratory of Bio-Resource and Eco-Environment of Ministry of Education, College of Life Sciences, Sichuan University, Chengdu 610064, China.
Organizational Affiliation: