Organic-Platinum Hybrids for Covalent Binding of G-Quadruplexes: Structural Basis and Application to Cancer Immunotherapy.
Liu, L.Y., Ma, T.Z., Zeng, Y.L., Liu, W., Zhang, H., Mao, Z.W.(2023) Angew Chem Int Ed Engl 62: e202305645-e202305645
- PubMed: 37464955 Search on PubMed
- DOI: https://doi.org/10.1002/anie.202305645
- Primary Citation Related Structures:
8IJC - PubMed Abstract:
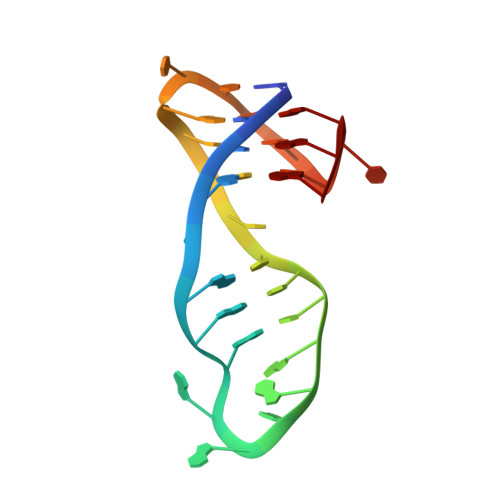
G-quadruplexes (G4s) have been revived as promising therapeutic targets with the development of immunotherapy, but the G4-mediated immune response remains unclear. We designed a novel class of G4-binding organic-platinum hybrids, L 1 -cispt and L 1 -transpt, with spatial matching for G4 binding and G4 DNA reactivity for binding site locking. The solution structure of L 1 -transpt-MYT1L G4 demonstrated the effectiveness of the covalent binding and revealed the covalent binding-guided dynamic balance, accompanied by the destruction of the A5-T17 base pairs to achieve the covalent binding of the platinum unit to N7 of the G6 residue. Furthermore, L 1 -cispt- and L 1 -transpt-mediated genomic dysfunction could activate the retinoic acid-induced gene I (RIG-I) pathway and induce immunogenic cell death (ICD). The use of L 1 -cispt/L 1 -transpt-treated dying cells as therapeutic vaccines stimulated a robust immune response and effectively inhibited tumor growth in vivo. Our findings highlight the importance of the rational combination of specific spatial recognition and covalent locking in G4-trageting drug design and their potential in immunotherapy.
- MOE Key Laboratory of Bioinorganic and Synthetic Chemistry, School of Chemistry, IGCME, Sun Yat-Sen University, Guangzhou, 510275, P. R. China.
Organizational Affiliation: