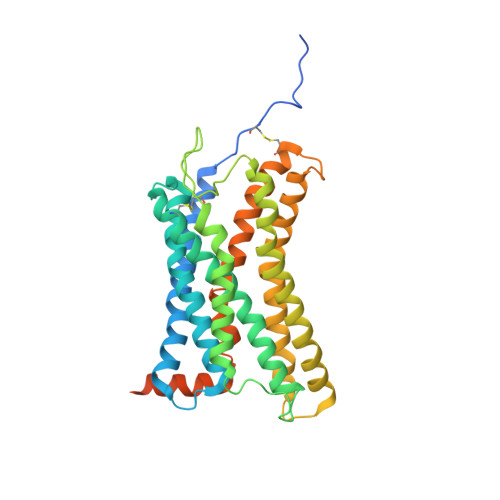
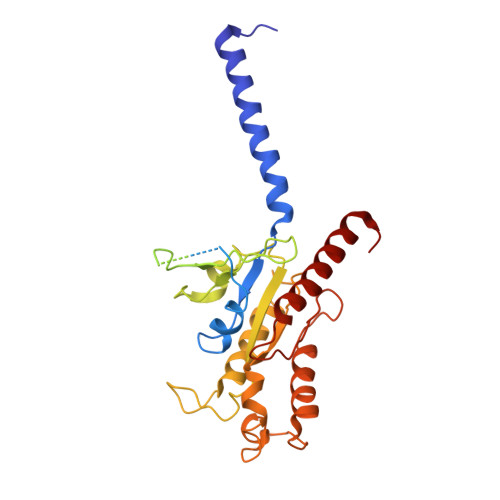
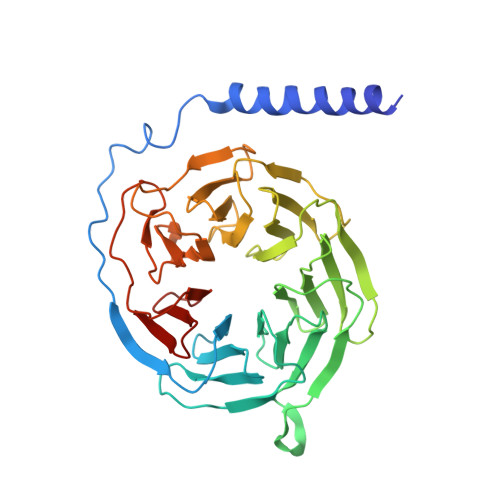
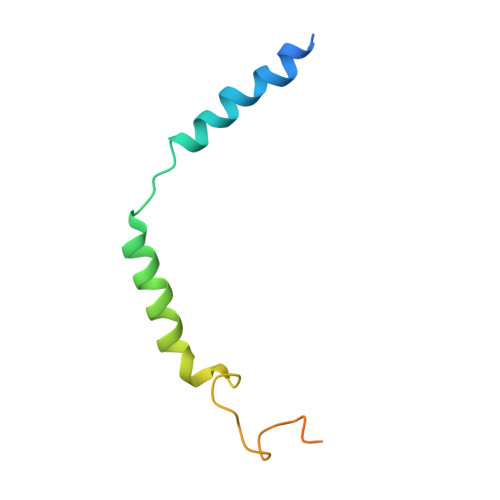
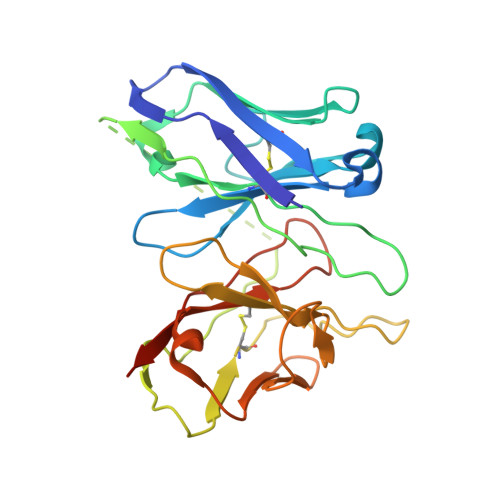
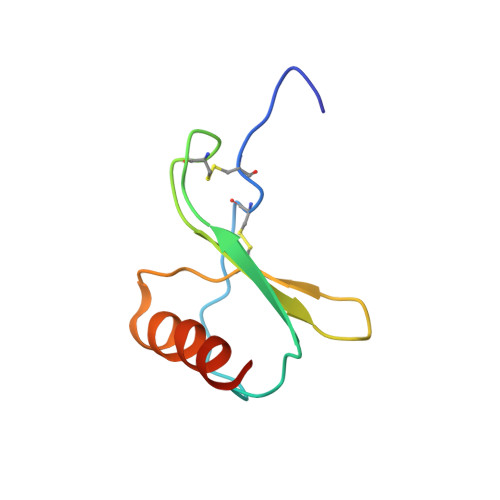
Structural basis of CXC chemokine receptor 1 ligand binding and activation.
Ishimoto, N., Park, J.H., Kawakami, K., Tajiri, M., Mizutani, K., Akashi, S., Tame, J.R.H., Inoue, A., Park, S.Y.(2023) Nat Commun 14: 4107-4107
- PubMed: 37433790 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-39799-2
- Primary Citation Related Structures:
8IC0 - PubMed Abstract:
Neutrophil granulocytes play key roles in innate immunity and shaping adaptive immune responses. They are attracted by chemokines to sites of infection and tissue damage, where they kill and phagocytose bacteria. The chemokine CXCL8 (also known as interleukin-8, abbreviated IL-8) and its G-protein-coupled receptors CXCR1 and CXCR2 are crucial elements in this process, and also the development of many cancers. These GPCRs have therefore been the target of many drug development campaigns and structural studies. Here, we solve the structure of CXCR1 complexed with CXCL8 and cognate G-proteins using cryo-EM, showing the detailed interactions between the receptor, the chemokine and Gαi protein. Unlike the closely related CXCR2, CXCR1 strongly prefers to bind CXCL8 in its monomeric form. The model shows that steric clashes would form between dimeric CXCL8 and extracellular loop 2 (ECL2) of CXCR1. Consistently, transplanting ECL2 of CXCR2 onto CXCR1 abolishes the selectivity for the monomeric chemokine. Our model and functional analysis of various CXCR1 mutants will assist efforts in structure-based drug design targeting specific CXC chemokine receptor subtypes.
- Drug Design Laboratory, Graduate School of Medical Life Science, Yokohama City University, Tsurumi, Yokohama, 230-0045, Japan.
Organizational Affiliation: