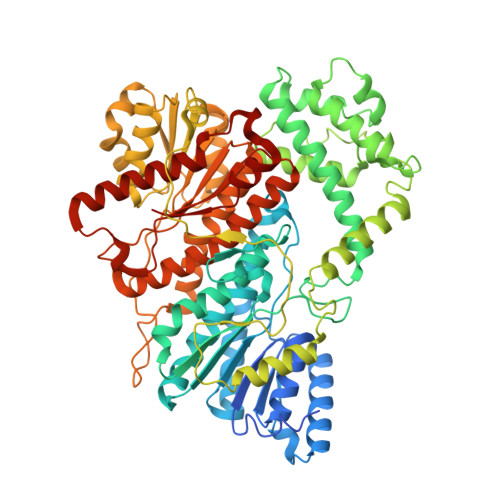
Cryo-EM structure of bifunctional malonyl-CoA reductase from Chloroflexus aurantiacus reveals a dynamic domain movement for high enzymatic activity.
Ahn, J.W., Kim, S., Hong, J., Kim, K.J.(2023) Int J Biol Macromol 242: 124676-124676
- PubMed: 37146856 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2023.124676
- Primary Citation Related Structures:
8HJW, 8I6Z, 8I70 - PubMed Abstract:
The platform chemical 3-hydroxypropionic acid is used to synthesize various valuable materials, including bioplastics. Bifunctional malonyl-CoA reductase is a key enzyme in 3-hydroxypropionic acid biosynthesis as it catalyzes the two-step reduction of malonyl-CoA to malonate semialdehyde to 3-hydroxypropionic acid. Here, we report the cryo-EM structure of a full-length malonyl-CoA reductase protein from Chloroflexus aurantiacus (CaMCR Full ). The EM model of CaMCR Full reveals a tandem helix architecture comprising an N-terminal (CaMCR ND ) and a C-terminal (CaMCR CD ) domain. The CaMCR Full model also revealed that the enzyme undergoes a dynamic domain movement between CaMCR ND and CaMCR CD due to the presence of a flexible linker between these two domains. Increasing the flexibility and extension of the linker resulted in a twofold increase in enzyme activity, indicating that for CaMCR, domain movement is crucial for high enzyme activity. We also describe the structural features of CaMCR ND and CaMCR CD . This study reveals the protein structures underlying the molecular mechanism of CaMCR Full and thereby provides valuable information for future enzyme engineering to improve the productivity of 3-hydroxypropionic acid.
- Postech Biotech Center, Pohang University of Science and Technology, 77 Cheongam-ro, Nam-gu, Pohang, Gyeongbuk 37673, Republic of Korea; Center for Biomolecular Capture Technology, Bio Open Innovation Center, Pohang University of Science and Technology, 47 Cheongam-ro, Nam-gu, Pohang, Gyeongbuk 37673, Republic of Korea.
Organizational Affiliation: