PGL-III, a Rare Intermediate of Mycobacterium leprae Phenolic Glycolipid Biosynthesis, Is a Potent Mincle Ligand.
Ishizuka, S., van Dijk, J.H.M., Kawakita, T., Miyamoto, Y., Maeda, Y., Goto, M., Le Calvez, G., Groot, L.M., Witte, M.D., Minnaard, A.J., van der Marel, G.A., Ato, M., Nagae, M., Codee, J.D.C., Yamasaki, S.(2023) ACS Cent Sci 9: 1388-1399
- PubMed: 37521780 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acscentsci.3c00040
- Primary Citation Related Structures:
8H4V, 8HB5 - PubMed Abstract:
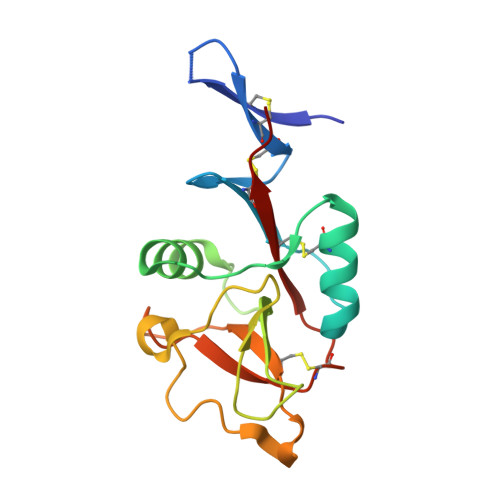
Although leprosy (Hansen's disease) is one of the oldest known diseases, the pathogenicity of Mycobacterium leprae ( M. leprae ) remains enigmatic. Indeed, the cell wall components responsible for the immune response against M. leprae are as yet largely unidentified. We reveal here phenolic glycolipid-III (PGL-III) as an M. leprae -specific ligand for the immune receptor Mincle. PGL-III is a scarcely present trisaccharide intermediate in the biosynthetic pathway to PGL-I, an abundant and characteristic M. leprae glycolipid. Using activity-based purification, we identified PGL-III as a Mincle ligand that is more potent than the well-known M. tuberculosis trehalose dimycolate. The cocrystal structure of Mincle and a synthetic PGL-III analogue revealed a unique recognition mode, implying that it can engage multiple Mincle molecules. In Mincle-deficient mice infected with M. leprae , increased bacterial burden with gross pathologies were observed. These results show that PGL-III is a noncanonical ligand recognized by Mincle, triggering protective immunity.
- Department of Molecular Immunology, Research Institute for Microbial Diseases, Osaka University, 3-1 Yamadaoka, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: