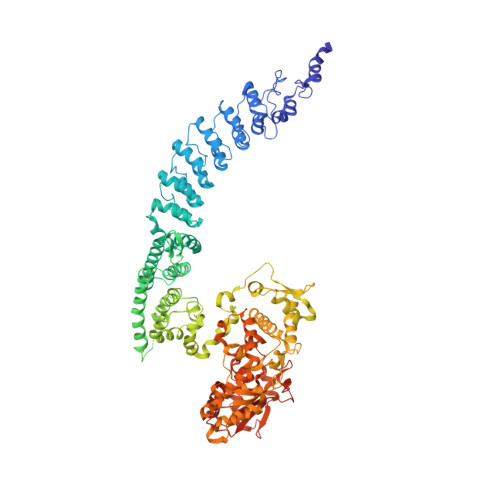
Structural Basis for the Enzymatic Activity of the HACE1 HECT-Type E3 Ligase Through N-Terminal Helix Dimerization.
Singh, S., Machida, S., Tulsian, N.K., Choong, Y.K., Ng, J., Shankar, S., Liu, Y., Chandiramani, K.V., Shi, J., Sivaraman, J.(2023) Adv Sci (Weinh) 10: e2207672-e2207672
- PubMed: 37537642 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/advs.202207672
- Primary Citation Related Structures:
8H8X, 8HAE - PubMed Abstract:
HACE1 is an ankyrin repeat (AKR) containing HECT-type E3 ubiquitin ligase that interacts with and ubiquitinates multiple substrates. While HACE1 is a well-known tumor suppressor, its structure and mode of ubiquitination are not understood. The authors present the cryo-EM structures of human HACE1 along with in vitro functional studies that provide insights into how the enzymatic activity of HACE1 is regulated. HACE1 comprises of an N-terminal AKR domain, a middle (MID) domain, and a C-terminal HECT domain. Its unique G-shaped architecture interacts as a homodimer, with monomers arranged in an antiparallel manner. In this dimeric arrangement, HACE1 ubiquitination activity is hampered, as the N-terminal helix of one monomer restricts access to the C-terminal domain of the other. The in vitro ubiquitination assays, hydrogen-deuterium exchange mass spectrometry (HDX-MS) analysis, mutagenesis, and in silico modeling suggest that the HACE1 MID domain plays a crucial role along with the AKRs in RAC1 substrate recognition.
- Department of Biological Sciences, National University of Singapore, 14 Science Drive 4, Singapore, 117558, Singapore.
Organizational Affiliation: