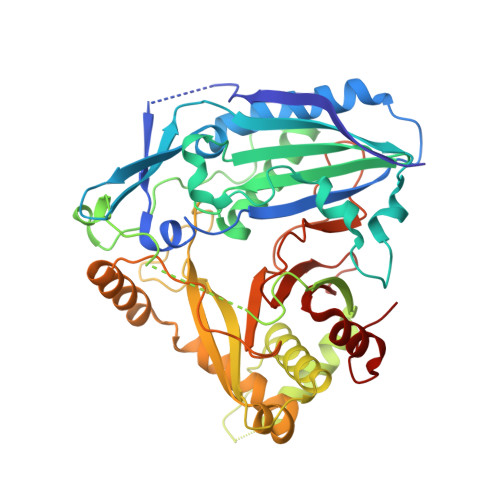
Characterization and structure-based protein engineering of a regiospecific saponin acetyltransferase from Astragalus membranaceus.
Wang, L., Jiang, Z., Zhang, J., Chen, K., Zhang, M., Wang, Z., Wang, B., Ye, M., Qiao, X.(2023) Nat Commun 14: 5969-5969
- PubMed: 37749089 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-41599-7
- Primary Citation Related Structures:
8H8I, 8HBT - PubMed Abstract:
Acetylation contributes to the bioactivity of numerous medicinally important natural products. However, little is known about the acetylation on sugar moieties. Here we report a saponin acetyltransferase from Astragalus membranaceus. AmAT7-3 is discovered through a stepwise gene mining approach and characterized as the xylose C3'/C4'-O-acetyltransferse of astragaloside IV (1). To elucidate its catalytic mechanism, complex crystal structures of AmAT7-3/1 and AmAT7-3 A310G /1 are obtained, which reveal a large active pocket decided by a specific sequence AADAG. Combining with QM/MM computation, the regiospecificity of AmAT7-3 is determined by sugar positioning modulated by surrounding amino acids including #A310 and #L290. Furthermore, a small mutant library is built using semi-rational design, where variants A310G and A310W are found to catalyze specific C3'-O and C4'-O acetylation, respectively. AmAT7-3 and its variants are also employed to acetylate other bioactive saponins. This work expands the understanding of saponin acetyltransferases, and provide efficient catalytic tools for saponin acetylation.
- State Key Laboratory of Natural and Biomimetic Drugs, School of Pharmaceutical Sciences, Peking University, 38 Xueyuan Road, Beijing, 100191, China.
Organizational Affiliation: