Cryo-EM structure of Escherichia coli cytochrome bo3 in DDM detergent
Cao, H.Y., Li, K., Li, C.Y.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cytochrome bo(3) ubiquinol oxidase subunit 1 | 663 | Escherichia coli K-12 | Mutation(s): 0 EC: 7.1.1.3 Membrane Entity: Yes | 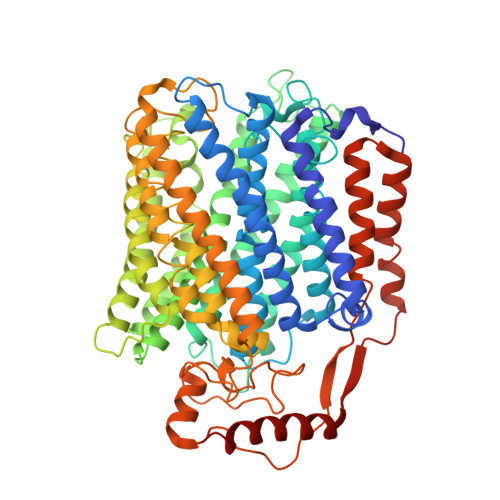 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0ABI8 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cytochrome bo(3) ubiquinol oxidase subunit 2 | 315 | Escherichia coli K-12 | Mutation(s): 0 Membrane Entity: Yes | 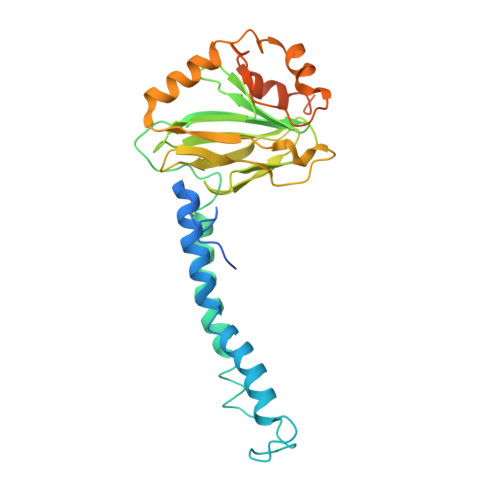 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0ABJ1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| ubiquinol oxidase | 204 | Escherichia coli K-12 | Mutation(s): 0 Membrane Entity: Yes | 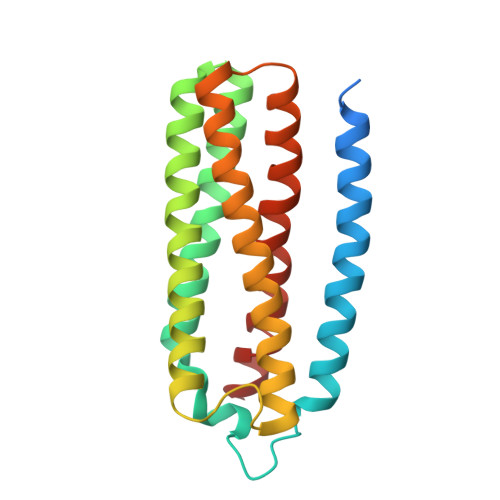 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0ABJ3 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cytochrome bo(3) ubiquinol oxidase subunit 4 | 109 | Escherichia coli K-12 | Mutation(s): 0 Membrane Entity: Yes | 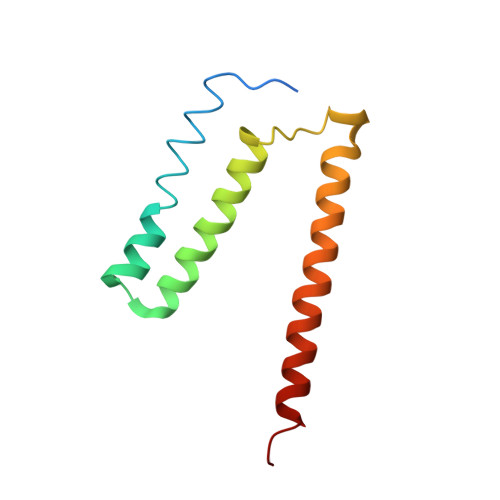 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0ABJ6 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| HEO Download:Ideal Coordinates CCD File | F [auth A] | HEME O C49 H58 Fe N4 O5 FRKORVCRVCLRBA-BZKSIRDQSA-L |  | ||
| 3PE Download:Ideal Coordinates CCD File | I [auth A], J [auth C] | 1,2-Distearoyl-sn-glycerophosphoethanolamine C41 H82 N O8 P LVNGJLRDBYCPGB-LDLOPFEMSA-N |  | ||
| UQ8 Download:Ideal Coordinates CCD File | H [auth A] | Ubiquinone-8 C49 H74 O4 ICFIZJQGJAJRSU-SGHXUWJISA-N |  | ||
| HEM Download:Ideal Coordinates CCD File | E [auth A] | PROTOPORPHYRIN IX CONTAINING FE C34 H32 Fe N4 O4 KABFMIBPWCXCRK-RGGAHWMASA-L |  | ||
| CU Download:Ideal Coordinates CCD File | G [auth A] | COPPER (II) ION Cu JPVYNHNXODAKFH-UHFFFAOYSA-N |  | ||
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |