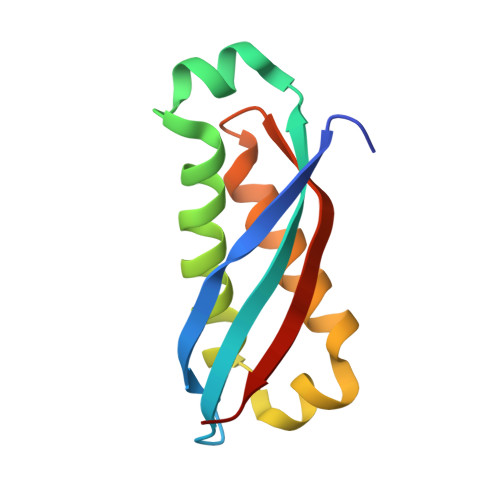
Structural and mutational studies suggest key residues to determine whether stomatin SPFH domains form dimers or trimers.
Komatsu, T., Matsui, I., Yokoyama, H.(2022) Biochem Biophys Rep 32: 101384-101384
- PubMed: 36386441 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbrep.2022.101384
- Primary Citation Related Structures:
8GN9 - PubMed Abstract:
Stomatin is a major integral membrane protein in human erythrocytes. In a form of hemolytic anemia known as hereditary stomatocytosis, stomatin is deficient in the erythrocyte membrane due to mis-trafficking. It is a member of stomatin, prohibitin, flotillin, and HflK/C (SPFH) domain proteins, and SPFH proteins could function as membrane-bound oligomeric scaffolding proteins in lipid rafts. The previously determined structure of the SPFH domain of Pyrococcus horikoshii (Ph) stomatin formed a trimer, whereas that of mouse stomatin formed a dimer. To elucidate the difference of oligomerization state, structural and chromatographic analyses using Ph stomatin were performed, and the key residues were suggested to determine whether SPFH domains form dimers or trimers. From gel-filtration analyses, PhStom (56-234) formed a trimer or tetramer, whereas PhStom (63-234) and PhStom (56-234) K59S formed a dimer. The residues 56-62, particularly Lys59, were involved in trimerization. Based on the crystal structure of PhStom (63-234), it formed a banana-shaped dimer, as observed in mouse stomatin. Thus, residues 162-168 are involved in dimerization. This study provides important insight into the molecular function and oligomerization state of stomatin.
- Faculty of Pharmaceutical Sciences, Tokyo University of Science, 2641 Yamazaki, Noda, Chiba, 278-8510, Japan.
Organizational Affiliation: