Single hormone or synthetic agonist induces G s /G i coupling selectivity of EP receptors via distinct binding modes and propagating paths.
Huang, S.M., Xiong, M.Y., Liu, L., Mu, J., Wang, M.W., Jia, Y.L., Cai, K., Tie, L., Zhang, C., Cao, S., Wen, X., Wang, J.L., Guo, S.C., Li, Y., Qu, C.X., He, Q.T., Cai, B.Y., Xue, C., Gan, S., Xie, Y., Cong, X., Yang, Z., Kong, W., Li, S., Li, Z., Xiao, P., Yang, F., Yu, X., Guan, Y.F., Zhang, X., Liu, Z., Yang, B.X., Du, Y., Sun, J.P.(2023) Proc Natl Acad Sci U S A 120: e2216329120-e2216329120
- PubMed: 37478163 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2216329120
- Primary Citation Related Structures:
8GCM, 8GCP, 8GD9, 8GDA, 8GDB, 8GDC - PubMed Abstract:
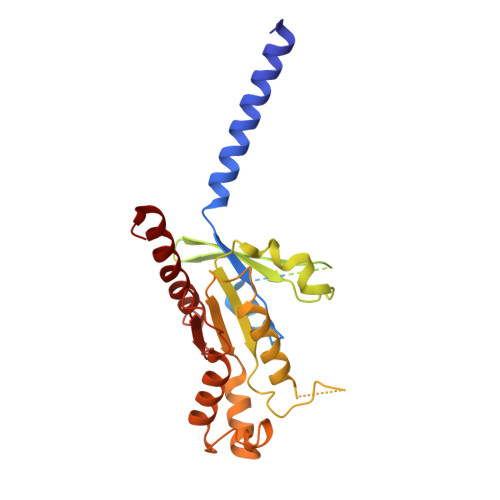
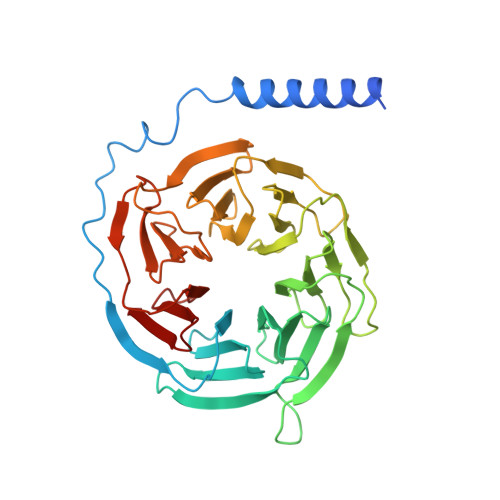
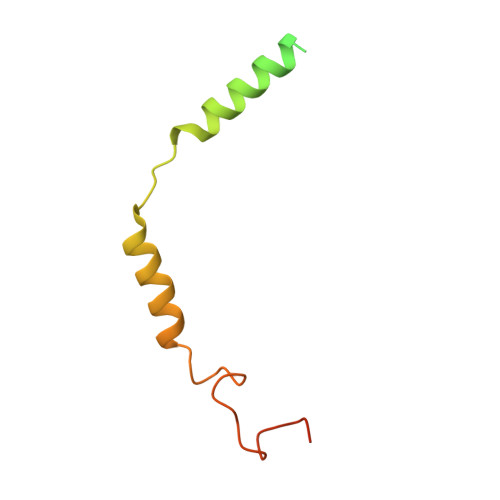
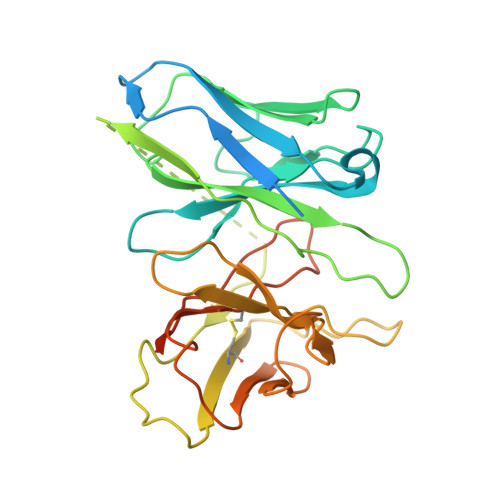
To accomplish concerted physiological reactions, nature has diversified functions of a single hormone at at least two primary levels: 1) Different receptors recognize the same hormone, and 2) different cellular effectors couple to the same hormone-receptor pair [R.P. Xiao, Sci STKE 2001 , re15 (2001); L. Hein, J. D. Altman, B.K. Kobilka, Nature 402 , 181-184 (1999); Y. Daaka, L. M. Luttrell, R. J. Lefkowitz, Nature 390 , 88-91 (1997)]. Not only these questions lie in the heart of hormone actions and receptor signaling but also dissecting mechanisms underlying these questions could offer therapeutic routes for refractory diseases, such as kidney injury (KI) or X-linked nephrogenic diabetes insipidus (NDI). Here, we identified that G s -biased signaling, but not G i activation downstream of EP4, showed beneficial effects for both KI and NDI treatments. Notably, by solving Cryo-electron microscope (cryo-EM) structures of EP3-G i , EP4-G s , and EP4-G i in complex with endogenous prostaglandin E 2 (PGE 2 )or two synthetic agonists and comparing with PGE 2 -EP2-G s structures, we found that unique primary sequences of prostaglandin E2 receptor (EP) receptors and distinct conformational states of the EP4 ligand pocket govern the G s /G i transducer coupling selectivity through different structural propagation paths, especially via TM6 and TM7, to generate selective cytoplasmic structural features. In particular, the orientation of the PGE 2 ω-chain and two distinct pockets encompassing agonist L902688 of EP4 were differentiated by their G s /G i coupling ability. Further, we identified common and distinct features of cytoplasmic side of EP receptors for G s /G i coupling and provide a structural basis for selective and biased agonist design of EP4 with therapeutic potential.
- Department of Physiology and Pathophysiology, School of Basic Medical Sciences, State Key Laboratory of Vascular Homeostasis and Remodeling, Peking University, Beijing 100191, China.
Organizational Affiliation: