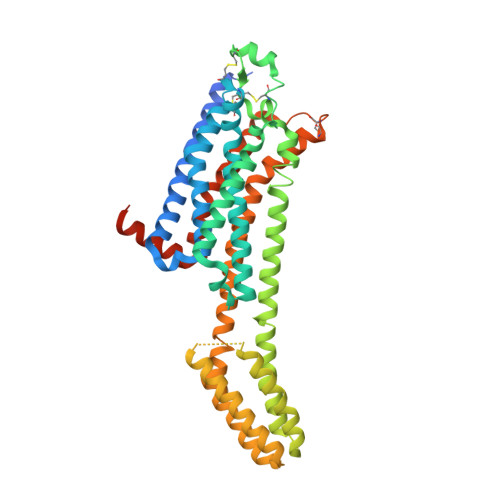
A robust approach for MicroED sample preparation of lipidic cubic phase embedded membrane protein crystals.
Martynowycz, M.W., Shiriaeva, A., Clabbers, M.T.B., Nicolas, W.J., Weaver, S.J., Hattne, J., Gonen, T.(2023) Nat Commun 14: 1086-1086
- PubMed: 36841804 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-36733-4
- Primary Citation Related Structures:
8FYN, 8FYO, 8FYP, 8FYQ, 8FYR, 8FYS - PubMed Abstract:
Crystallizing G protein-coupled receptors (GPCRs) in lipidic cubic phase (LCP) often yields crystals suited for the cryogenic electron microscopy (cryoEM) method microcrystal electron diffraction (MicroED). However, sample preparation is challenging. Embedded crystals cannot be targeted topologically. Here, we use an integrated fluorescence light microscope (iFLM) inside of a focused ion beam and scanning electron microscope (FIB-SEM) to identify fluorescently labeled GPCR crystals. Crystals are targeted using the iFLM and LCP is milled using a plasma focused ion beam (pFIB). The optimal ion source for preparing biological lamellae is identified using standard crystals of proteinase K. Lamellae prepared using either argon or xenon produced the highest quality data and structures. MicroED data are collected from the milled lamellae and the structures are determined. This study outlines a robust approach to identify and mill membrane protein crystals for MicroED and demonstrates plasma ion-beam milling is a powerful tool for preparing biological lamellae.
- Howard Hughes Medical Institute, University of California, Los Angeles, CA, 90095, USA.
Organizational Affiliation: