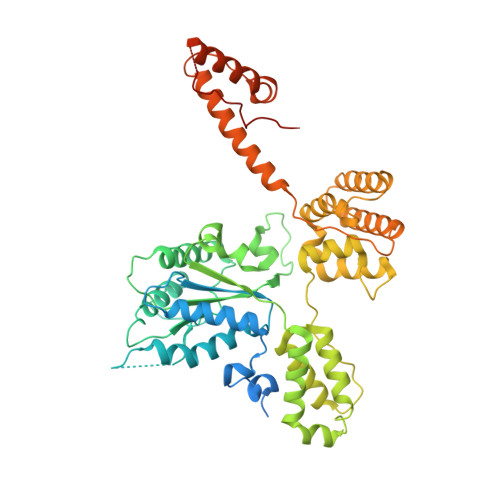
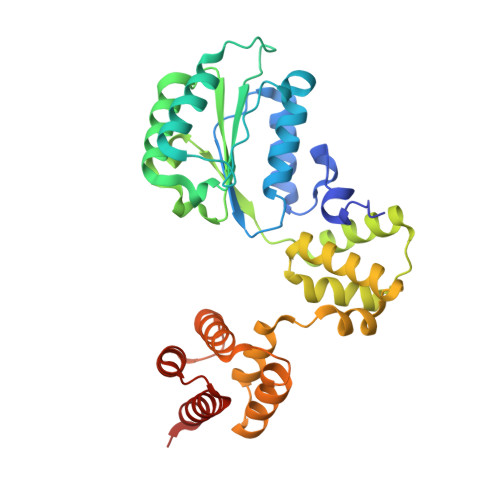
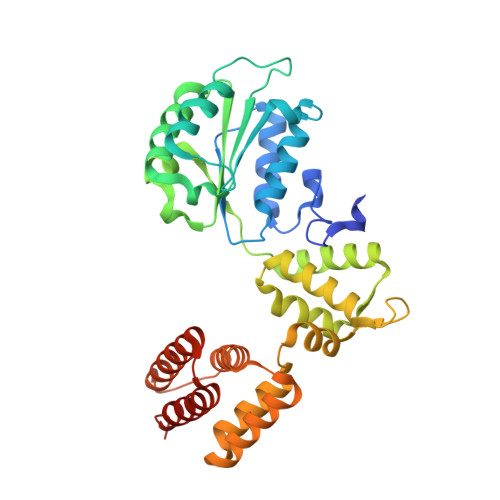
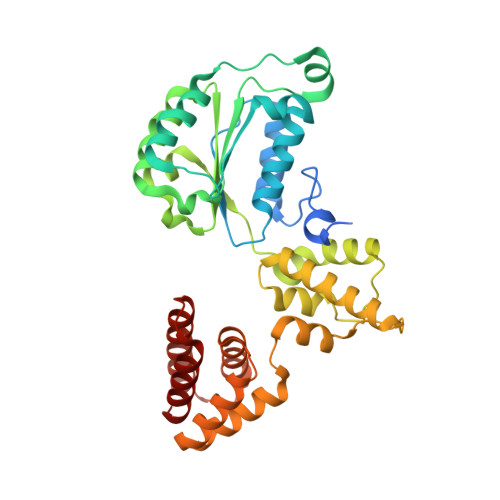
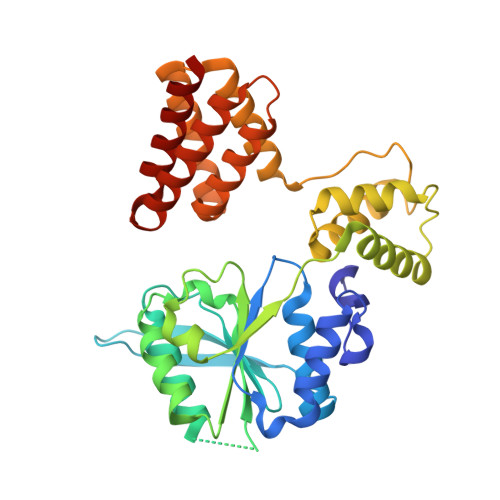
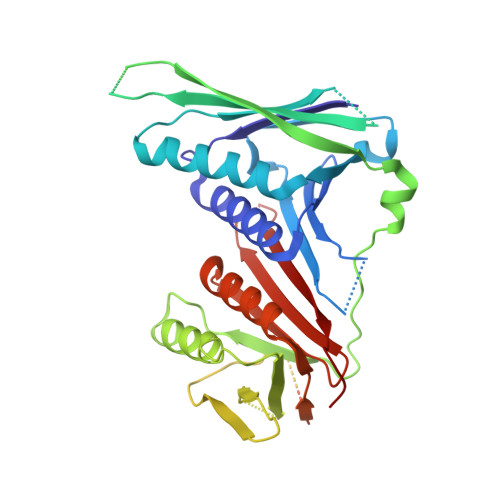
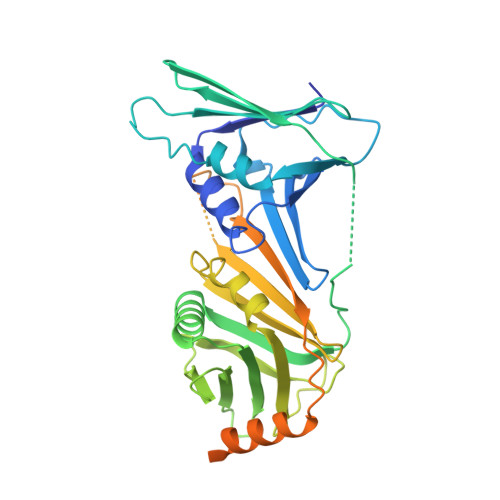
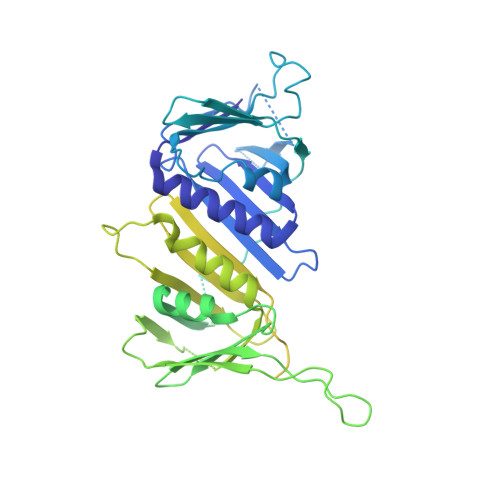
Structures of 9-1-1 DNA checkpoint clamp loading at gaps from start to finish and ramification on biology.
Zheng, F., Georgescu, R.E., Yao, N.Y., O'Donnell, M.E., Li, H.(2023) Cell Rep 42: 112694-112694
- PubMed: 37392384 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2023.112694
- Primary Citation Related Structures:
8FS3, 8FS4, 8FS5, 8FS6, 8FS7, 8FS8 - PubMed Abstract:
Rad24-RFC (replication factor C) loads the 9-1-1 checkpoint clamp onto the recessed 5' ends by binding a 5' DNA at an external surface site and threading the 3' single-stranded DNA (ssDNA) into 9-1-1. We find here that Rad24-RFC loads 9-1-1 onto DNA gaps in preference to a recessed 5' end, thus presumably leaving 9-1-1 on duplex 3' ss/double-stranded DNA (dsDNA) after Rad24-RFC ejects from DNA. We captured five Rad24-RFC-9-1-1 loading intermediates using a 10-nt gap DNA. We also determined the structure of Rad24-RFC-9-1-1 using a 5-nt gap DNA. The structures reveal that Rad24-RFC is unable to melt DNA ends and that a Rad24 loop limits the dsDNA length in the chamber. These observations explain Rad24-RFC's preference for a preexisting gap of over 5-nt ssDNA and suggest a direct role of the 9-1-1 in gap repair with various TLS (trans-lesion synthesis) polymerases in addition to signaling the ATR kinase.
- Department of Structural Biology, Van Andel Institute, Grand Rapids, MI, USA.
Organizational Affiliation: