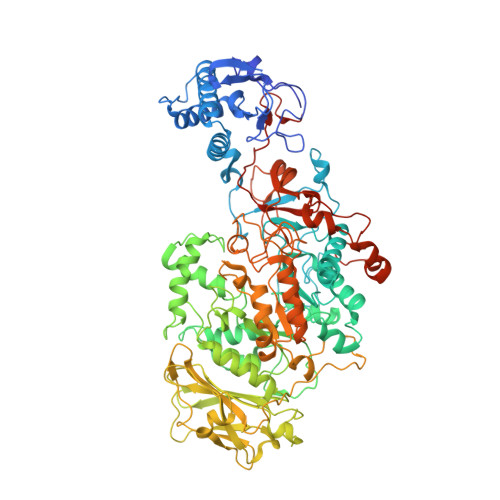
The catalytic domains of Streptococcus mutans glucosyltransferases: a structural analysis.
Schormann, N., Patel, M., Thannickal, L., Purushotham, S., Wu, R., Mieher, J.L., Wu, H., Deivanayagam, C.(2023) Acta Crystallogr F Struct Biol Commun 79: 119-127
- PubMed: 37158310 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X23003199
- Primary Citation Related Structures:
8FJ9, 8FJC, 8FK4, 8FKL, 8FN5 - PubMed Abstract:
Streptococcus mutans, found in the human oral cavity, is a significant contributor to the pathogenesis of dental caries. This bacterium expresses three genetically distinct types of glucosyltransferases named GtfB (GTF-I), GtfC (GTF-SI) and GtfD (GTF-S) that play critical roles in the development of dental plaque. The catalytic domains of GtfB, GtfC and GtfD contain conserved active-site residues for the overall enzymatic activity that relate to hydrolytic glycosidic cleavage of sucrose to glucose and fructose, release of fructose and generation of a glycosyl-enzyme intermediate in the reducing end. In a subsequent transglycosylation step, the glucosyl moiety is transferred to the nonreducing end of an acceptor to form a growing glucan polymer chain made up of glucose molecules. It has been proposed that both sucrose breakdown and glucan synthesis occur in the same active site of the catalytic domain, although the active site does not appear to be large enough to accommodate both functions. These three enzymes belong to glycoside hydrolase family 70 (GH70), which shows homology to glycoside hydrolase family 13 (GH13). GtfC synthesizes both soluble and insoluble glucans (α-1,3 and α-1,6 glycosidic linkages), while GtfB and GtfD synthesize only insoluble or soluble glucans, respectively. Here, crystal structures of the catalytic domains of GtfB and GtfD are reported. These structures are compared with previously determined structures of the catalytic domain of GtfC. With this work, apo structures and inhibitor-complex structures with acarbose are now available for the catalytic domains of GtfC and GtfB. The structure of GtfC with maltose allows further identification and comparison of active-site residues. A model of sucrose binding to GtfB is also included. The new structure of the catalytic domain of GtfD affords a structural comparison of the three S. mutans glycosyltransferases. Unfortunately, the catalytic domain of GtfD is not complete since crystallization resulted in the structure of a truncated protein lacking approximately 200 N-terminal residues of domain IV.
- Department of Biochemistry and Molecular Genetics, University of Alabama at Birmingham, Birmingham, AL 35294, USA.
Organizational Affiliation: