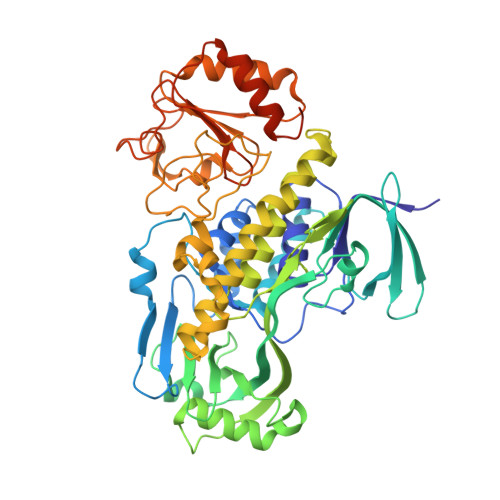
Crystal structure of MbnF: an NADPH-dependent flavin monooxygenase from Methylocystis strain SB2.
Stewart, A., Dershwitz, P., Stewart Jr., C., Sawaya, M.R., Yeates, T.O., Semrau, J.D., Zischka, H., DiSpirito, A.A., Bobik, T.A.(2023) Acta Crystallogr F Struct Biol Commun 79: 111-118
- PubMed: 37158309 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X23003035
- Primary Citation Related Structures:
8FHJ - PubMed Abstract:
Methanobactins (MBs) are ribosomally produced and post-translationally modified peptides (RiPPs) that are used by methanotrophs for copper acquisition. The signature post-translational modification of MBs is the formation of two heterocyclic groups, either an oxazolone, pyrazinedione or imidazolone group, with an associated thioamide from an X-Cys dipeptide. The precursor peptide (MbnA) for MB formation is found in a gene cluster of MB-associated genes. The exact biosynthetic pathway of MB formation is not yet fully understood, and there are still uncharacterized proteins in some MB gene clusters, particularly those that produce pyrazinedione or imidazolone rings. One such protein is MbnF, which is proposed to be a flavin monooxygenase (FMO) based on homology. To help to elucidate its possible function, MbnF from Methylocystis sp. strain SB2 was recombinantly produced in Escherichia coli and its X-ray crystal structure was resolved to 2.6 Å resolution. Based on its structural features, MbnF appears to be a type A FMO, most of which catalyze hydroxylation reactions. Preliminary functional characterization shows that MbnF preferentially oxidizes NADPH over NADH, supporting NAD(P)H-mediated flavin reduction, which is the initial step in the reaction cycle of several type A FMO enzymes. It is also shown that MbnF binds the precursor peptide for MB, with subsequent loss of the leader peptide sequence as well as the last three C-terminal amino acids, suggesting that MbnF might be needed for this process to occur. Finally, molecular-dynamics simulations revealed a channel in MbnF that is capable of accommodating the core MbnA fragment minus the three C-terminal amino acids.
- Roy J. Carver Department of Biochemistry, Biophysics and Molecular Biology, Iowa State University, Ames, IA 50011-3260, USA.
Organizational Affiliation: