Epitope topography of agonist antibodies to the checkpoint inhibitory receptor BTLA.
Cheung, T.C., Atwell, S., Bafetti, L., Cuenca, P.D., Froning, K., Hendle, J., Hickey, M., Ho, C., Huang, J., Lieu, R., Lim, S., Lippner, D., Obungu, V., Ward-Kavanagh, L., Weichert, K., Ware, C.F., Vendel, A.C.(2023) Structure 31: 958
- PubMed: 37279757 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2023.05.011
- Primary Citation Related Structures:
8F60, 8F6L, 8F6O - PubMed Abstract:
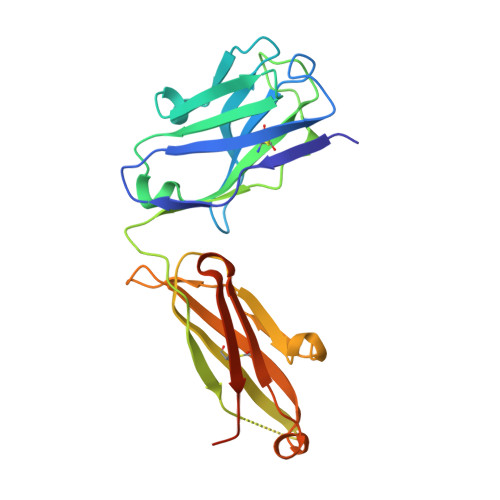
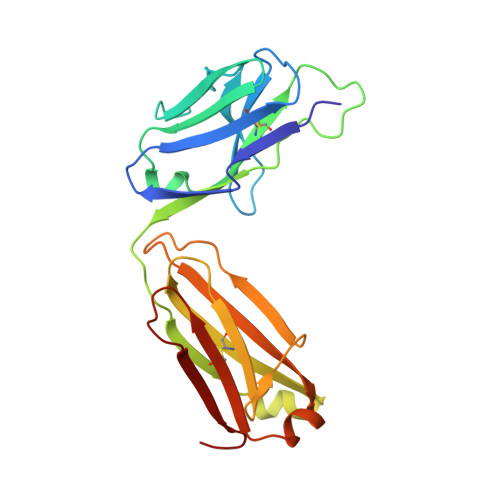
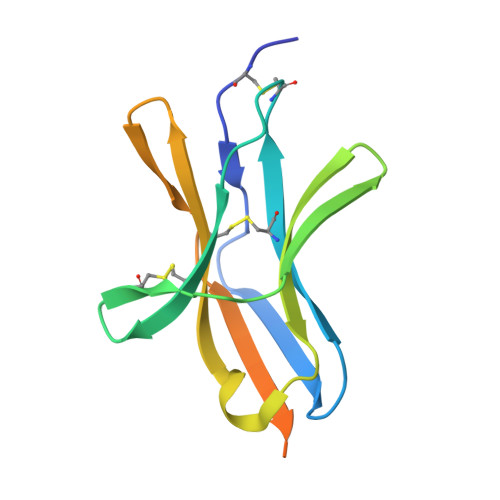
B and T lymphocyte attenuator (BTLA) is an attractive target for a new class of therapeutics that attempt to rebalance the immune system by agonizing checkpoint inhibitory receptors (CIRs). Herpesvirus entry mediator (HVEM) binds BTLA in both trans- and cis-orientations. We report here the development and structural characterization of three humanized BTLA agonist antibodies, 22B3, 25F7, and 23C8. We determined the crystal structures of the antibody-BTLA complexes, showing that these antibodies bind distinct and non-overlapping epitopes of BTLA. While all three antibodies activate BTLA, 22B3 mimics HVEM binding to BTLA and shows the strongest agonistic activity in functional cell assays and in an imiquimod-induced mouse model of psoriasis. 22B3 is also capable of modulating HVEM signaling through the BTLA-HVEM cis-interaction. The data obtained from crystal structures, biochemical assays, and functional studies provide a mechanistic model of HVEM and BTLA organization on the cell surface and informed the discovery of a highly active BTLA agonist.
- Infectious and Inflammatory Diseases Center, Sanford Burnham Prebys Medical Discovery Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: