Antibody Targeting of Conserved Sites of Vulnerability on the SARS-CoV-2 Spike Receptor-Binding Domain
Sankhala, R.S., Jensen, J., Modjarrad, K., Nelson, M., Joyce, M.G.To be published.
Experimental Data Snapshot
Starting Models: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Spike protein S1 | 205 | Severe acute respiratory syndrome coronavirus 2 | Mutation(s): 0 Gene Names: S, 2 | 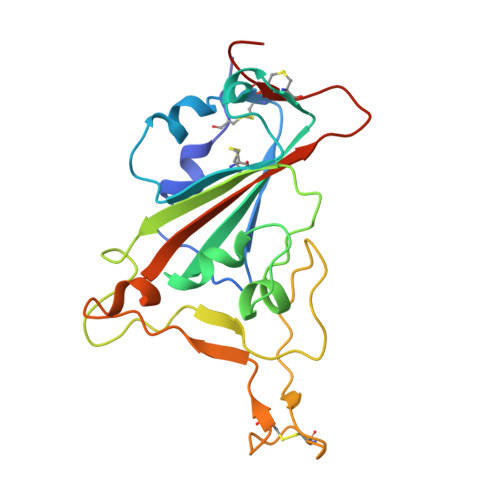 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0DTC2 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: P0DTC2-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| WRAIR-2123 Fab Heavy chain | B [auth H] | 232 | Homo sapiens | Mutation(s): 0 | 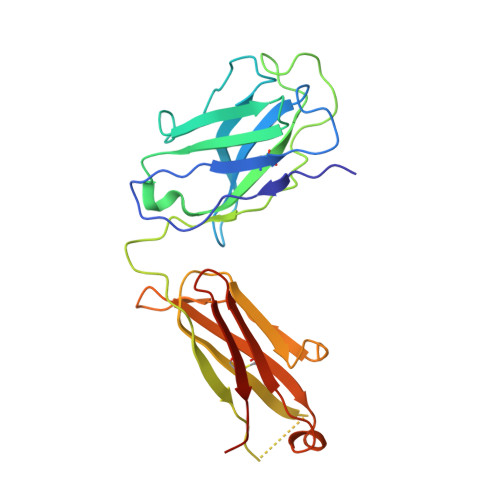 |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| WRAIR-2123 Fab Light chain | C [auth L] | 214 | Homo sapiens | Mutation(s): 0 | 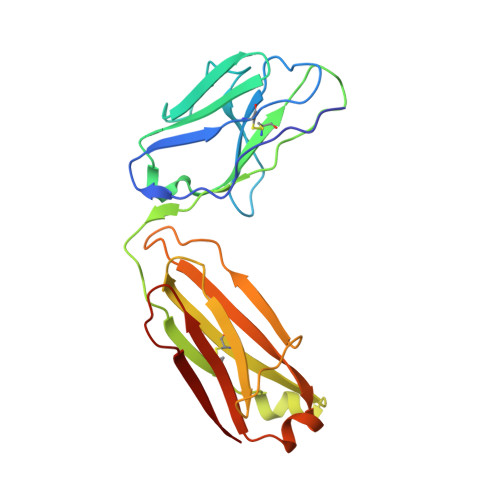 |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 39.29 | α = 90 |
| b = 94.43 | β = 90 |
| c = 201.31 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Department of Defense (DOD, United States) | United States | W81XWH-07-2-0067 |