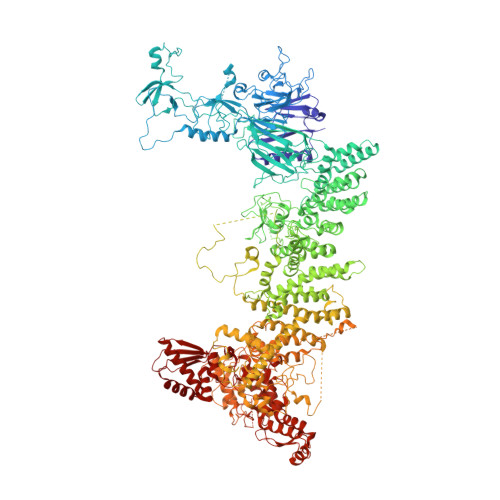
Structure of the human UBR5 E3 ubiquitin ligase.
Wang, F., He, Q., Zhan, W., Yu, Z., Finkin-Groner, E., Ma, X., Lin, G., Li, H.(2023) Structure 31: 541-552.e4
- PubMed: 37040767 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2023.03.010
- Primary Citation Related Structures:
8D4X, 8E0Q, 8EWI - PubMed Abstract:
The human UBR5 is a single polypeptide chain homology to E6AP C terminus (HECT)-type E3 ubiquitin ligase essential for embryonic development in mammals. Dysregulated UBR5 functions like an oncoprotein to promote cancer growth and metastasis. Here, we report that UBR5 assembles into a dimer and a tetramer. Our cryoelectron microscopy (cryo-EM) structures reveal that two crescent-shaped UBR5 monomers assemble head to tail to form the dimer, and two dimers bind face to face to form the cage-like tetramer with all four catalytic HECT domains facing the central cavity. Importantly, the N-terminal region of one subunit and the HECT of the other form an "intermolecular jaw" in the dimer. We show the jaw-lining residues are important for function, suggesting that the intermolecular jaw functions to recruit ubiquitin-loaded E2 to UBR5. Further work is needed to understand how oligomerization regulates UBR5 ligase activity. This work provides a framework for structure-based anticancer drug development and contributes to a growing appreciation of E3 ligase diversity.
- Department of Structural Biology, Van Andel Institute, 333 Bostwick Avenue NE, Grand Rapids, MI 49503, USA.
Organizational Affiliation: