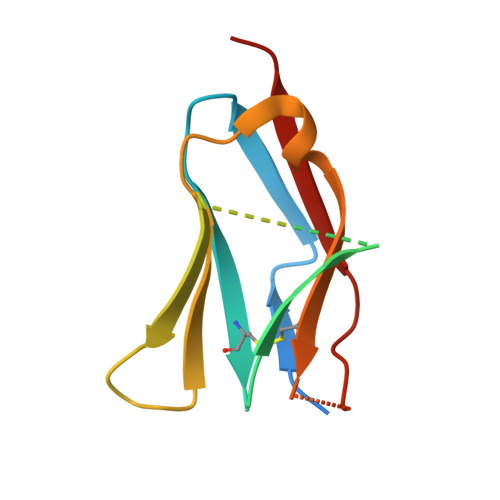
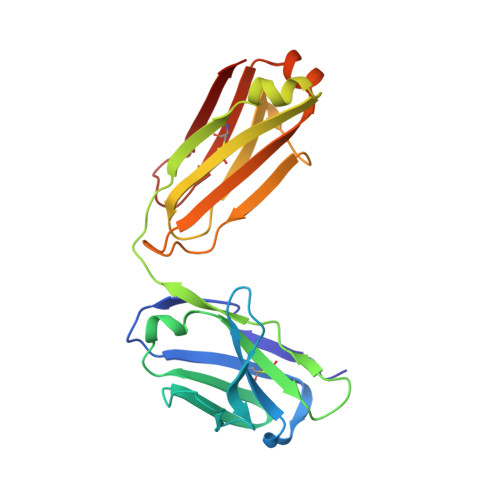
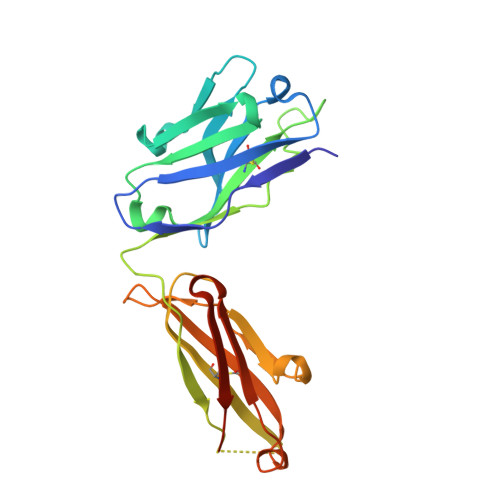
Antibody agonists trigger immune receptor signaling through local exclusion of receptor-type protein tyrosine phosphatases.
Lippert, A.H., Paluch, C., Gaglioni, M., Vuong, M.T., McColl, J., Jenkins, E., Fellermeyer, M., Clarke, J., Sharma, S., Moreira da Silva, S., Akkaya, B., Anzilotti, C., Morgan, S.H., Jessup, C.F., Korbel, M., Gileadi, U., Leitner, J., Knox, R., Chirifu, M., Huo, J., Yu, S., Ashman, N., Lui, Y., Wilkinson, I., Attfield, K.E., Fugger, L., Robertson, N.J., Lynch, C.J., Murray, L., Steinberger, P., Santos, A.M., Lee, S.F., Cornall, R.J., Klenerman, D., Davis, S.J.(2024) Immunity 57: 256-270.e10
- PubMed: 38354703 Search on PubMed
- DOI: https://doi.org/10.1016/j.immuni.2024.01.007
- Primary Citation Related Structures:
8EQ6, 9HK1 - PubMed Abstract:
Antibodies can block immune receptor engagement or trigger the receptor machinery to initiate signaling. We hypothesized that antibody agonists trigger signaling by sterically excluding large receptor-type protein tyrosine phosphatases (RPTPs) such as CD45 from sites of receptor engagement. An agonist targeting the costimulatory receptor CD28 produced signals that depended on antibody immobilization and were sensitive to the sizes of the receptor, the RPTPs, and the antibody itself. Although both the agonist and a non-agonistic anti-CD28 antibody locally excluded CD45, the agonistic antibody was more effective. An anti-PD-1 antibody that bound membrane proximally excluded CD45, triggered Src homology 2 domain-containing phosphatase 2 recruitment, and suppressed systemic lupus erythematosus and delayed-type hypersensitivity in experimental models. Paradoxically, nivolumab and pembrolizumab, anti-PD-1-blocking antibodies used clinically, also excluded CD45 and were agonistic in certain settings. Reducing these agonistic effects using antibody engineering improved PD-1 blockade. These findings establish a framework for developing new and improved therapies for autoimmunity and cancer.
- Department of Chemistry, University of Cambridge, Cambridge, UK.
Organizational Affiliation: