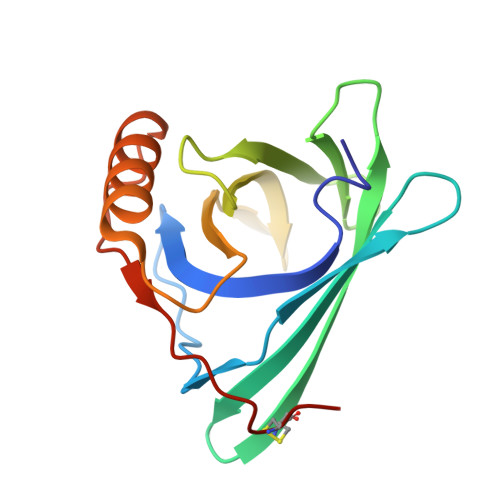
Structural and ligand binding analysis of the pet allergens Can f 1 and Fel d 7.
Min, J., Foo, A.C.Y., Gabel, S.A., Perera, L., DeRose, E.F., Pomes, A., Pedersen, L.C., Mueller, G.A.(2023) Front Allergy 4: 1133412-1133412
- PubMed: 36960093 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/falgy.2023.1133412
- Primary Citation Related Structures:
8EPU, 8EPV - PubMed Abstract:
Pet lipocalins are respiratory allergens with a central hydrophobic ligand-binding cavity called a calyx. Molecules carried in the calyx by allergens are suggested to influence allergenicity, but little is known about the native ligands. To provide more information on prospective ligands, we report crystal structures, NMR, molecular dynamics, and florescence studies of a dog lipocalin allergen Can f 1 and its closely related (and cross-reactive) cat allergen Fel d 7. Structural comparisons with reported lipocalins revealed that Can f 1 and Fel d 7 calyxes are open and positively charged while other dog lipocalin allergens are closed and negatively charged. We screened fatty acids as surrogate ligands, and found that Can f 1 and Fel d 7 bind multiple ligands with preferences for palmitic acid (16:0) among saturated fatty acids and oleic acid (18:1 cis-9) among unsaturated ones. NMR analysis of methyl probes reveals that conformational changes occur upon binding of pinolenic acid inside the calyx. Molecular dynamics simulation shows that the carboxylic group of fatty acids shuttles between two positively charged amino acids inside the Can f 1 and Fel d 7 calyx. Consistent with simulations, the stoichiometry of oleic acid-binding is 2:1 (fatty acid: protein) for Can f 1 and Fel d 7. The results provide valuable insights into the determinants of selectivity and candidate ligands for pet lipocalin allergens Can f 1 and Fel d 7.
- Genome Integrity and Structural Biology Laboratory, National Institute of Environmental Health Sciences, Durham, NC, United States.
Organizational Affiliation: