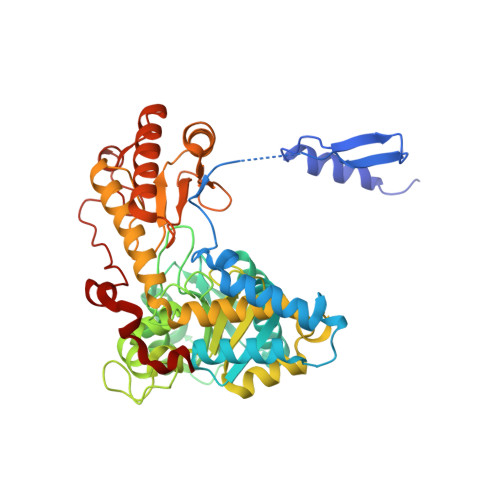
The yeast ALA synthase C-terminus positively controls enzyme structure and function.
Tran, J.U., Brown, B.L.(2023) Protein Sci 32: e4600-e4600
- PubMed: 36807942 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.4600
- Primary Citation Related Structures:
8EIM - PubMed Abstract:
5-Aminolevulinic acid synthase (ALAS) is a pyridoxal 5'-phosphate (PLP)-dependent enzyme that catalyzes the first and rate-limiting step of heme biosynthesis in α-proteobacteria and several non-plant eukaryotes. All ALAS homologs contain a highly conserved catalytic core, but eukaryotes also have a unique C-terminal extension that plays a role in enzyme regulation. Several mutations in this region are implicated in multiple blood disorders in humans. In Saccharomyces cerevisiae ALAS (Hem1), the C-terminal extension wraps around the homodimer core to contact conserved ALAS motifs proximal to the opposite active site. To determine the importance of these Hem1 C-terminal interactions, we determined the crystal structure of S. cerevisiae Hem1 lacking the terminal 14 amino acids (Hem1 ΔCT). With truncation of the C-terminal extension, we show structurally and biochemically that multiple catalytic motifs become flexible, including an antiparallel β-sheet important to Fold-Type I PLP-dependent enzymes. The changes in protein conformation result in an altered cofactor microenvironment, decreased enzyme activity and catalytic efficiency, and ablation of subunit cooperativity. These findings suggest that the eukaryotic ALAS C-terminus has a homolog-specific role in mediating heme biosynthesis, indicating a mechanism for autoregulation that can be exploited to allosterically modulate heme biosynthesis in different organisms.
- Department of Biochemistry, Vanderbilt University School of Medicine, Nashville, Tennessee, USA.
Organizational Affiliation: