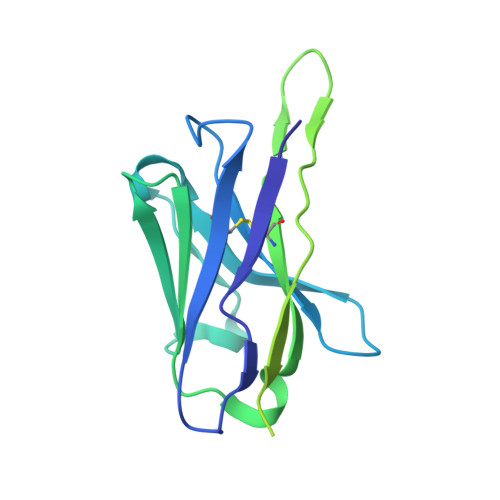
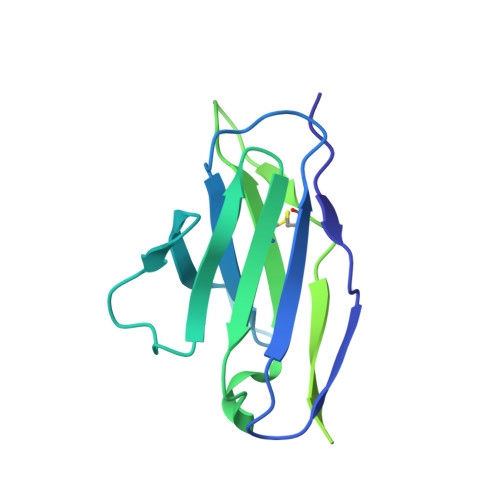
Structural basis of epitope selectivity and potent protection from malaria by PfCSP antibody L9.
Martin, G.M., Fernandez-Quintero, M.L., Lee, W.H., Pholcharee, T., Eshun-Wilson, L., Liedl, K.R., Pancera, M., Seder, R.A., Wilson, I.A., Ward, A.B.(2023) Nat Commun 14: 2815-2815
- PubMed: 37198165 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-38509-2
- Primary Citation Related Structures:
8EH5 - PubMed Abstract:
A primary objective in malaria vaccine design is the generation of high-quality antibody responses against the circumsporozoite protein of the malaria parasite, Plasmodium falciparum (PfCSP). To enable rational antigen design, we solved a cryo-EM structure of the highly potent anti-PfCSP antibody L9 in complex with recombinant PfCSP. We found that L9 Fab binds multivalently to the minor (NPNV) repeat domain, which is stabilized by a unique set of affinity-matured homotypic, antibody-antibody contacts. Molecular dynamics simulations revealed a critical role of the L9 light chain in integrity of the homotypic interface, which likely impacts PfCSP affinity and protective efficacy. These findings reveal the molecular mechanism of the unique NPNV selectivity of L9 and emphasize the importance of anti-homotypic affinity maturation in protective immunity against P. falciparum.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA, 92037, USA.
Organizational Affiliation: