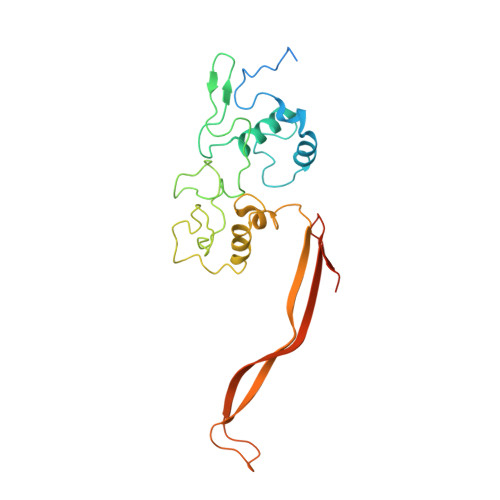
Extracellular cytochrome nanowires appear to be ubiquitous in prokaryotes.
Baquero, D.P., Cvirkaite-Krupovic, V., Hu, S.S., Fields, J.L., Liu, X., Rensing, C., Egelman, E.H., Krupovic, M., Wang, F.(2023) Cell 186: 2853-2864.e8
- PubMed: 37290436 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2023.05.012
- Primary Citation Related Structures:
8E5F, 8E5G - PubMed Abstract:
Electrically conductive appendages from the anaerobic bacterium Geobacter sulfurreducens, recently identified as extracellular cytochrome nanowires (ECNs), have received wide attention due to numerous potential applications. However, whether other organisms employ similar ECNs for electron transfer remains unknown. Here, using cryoelectron microscopy, we describe the atomic structures of two ECNs from two major orders of hyperthermophilic archaea present in deep-sea hydrothermal vents and terrestrial hot springs. Homologs of Archaeoglobus veneficus ECN are widespread among mesophilic methane-oxidizing Methanoperedenaceae, alkane-degrading Syntrophoarchaeales archaea, and in the recently described megaplasmids called Borgs. The ECN protein subunits lack similarities in their folds; however, they share a common heme arrangement, suggesting an evolutionarily optimized heme packing for efficient electron transfer. The detection of ECNs in archaea suggests that filaments containing closely stacked hemes may be a common and widespread mechanism for long-range electron transfer in both prokaryotic domains of life.
- Institut Pasteur, Université Paris Cité, CNRS UMR6047, Archaeal Virology Unit, Paris 75015, France.
Organizational Affiliation: