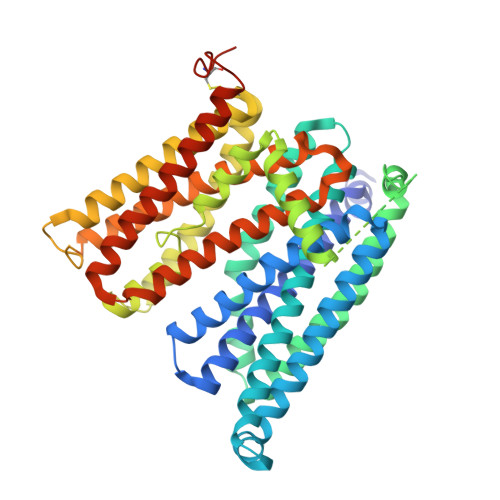
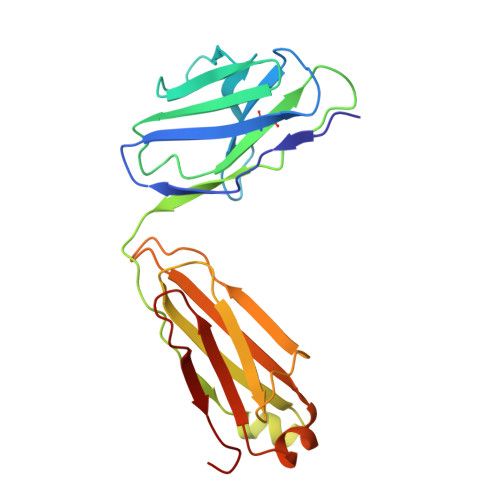
Mechanism of Ca 2+ transport by ferroportin.
Shen, J., Wilbon, A.S., Zhou, M., Pan, Y.(2023) Elife 12
- PubMed: 36648329 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.82947
- Primary Citation Related Structures:
8DL6 - PubMed Abstract:
Ferroportin (Fpn) is a transporter that releases ferrous ion (Fe 2+ ) from cells and is important for homeostasis of iron in circulation. Export of one Fe 2+ by Fpn is coupled to import of two H + to maintain charge balance. Here, we show that human Fpn (HsFpn) binds to and mediates Ca 2+ transport. We determine the structure of Ca 2+ -bound HsFpn and identify a single Ca 2+ binding site distinct from the Fe 2+ binding sites. Further studies validate the Ca 2+ binding site and show that Ca 2+ transport is not coupled to transport of another ion. In addition, Ca 2+ transport is significantly inhibited in the presence of Fe 2+ but not vice versa. Function of Fpn as a Ca 2+ uniporter may allow regulation of iron homeostasis by Ca 2+ .
- Verna and Marrs McLean Department of Biochemistry and Molecular Biology, Baylor College of Medicine, Houston, United States.
Organizational Affiliation: