S. CEREVISIAE CYP51 COMPLEXED WITH Courmarin-containing INHIBITOR
Benhamou, R., Keniya, M.V., Ruma, Y.N., Fridman, M., Tyndall, J.D., Monk, B.C.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Lanosterol 14-alpha demethylase | 539 | Saccharomyces cerevisiae YJM789 | Mutation(s): 0 Gene Names: ERG11, SCY_2394 EC: 1.14.14.154 Membrane Entity: Yes | 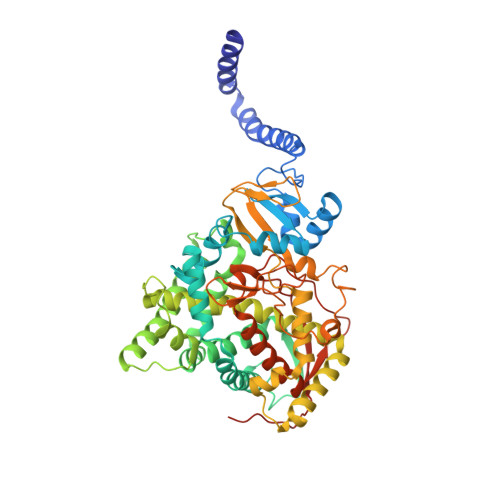 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P10614 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| HEM Download:Ideal Coordinates CCD File | B [auth A] | PROTOPORPHYRIN IX CONTAINING FE C34 H32 Fe N4 O4 KABFMIBPWCXCRK-RGGAHWMASA-L |  | ||
| SKX (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth A] | 7-(diethylamino)-N-[(2S)-2-(2,4-difluorophenyl)-2-hydroxy-3-(1H-1,2,4-triazol-1-yl)propyl]-2-oxo-2H-1-benzopyran-3-carboxamide C25 H25 F2 N5 O4 QYBYHULZZSKXIJ-VWLOTQADSA-N |  | ||
| 1PE Download:Ideal Coordinates CCD File | D [auth A] | PENTAETHYLENE GLYCOL C10 H22 O6 JLFNLZLINWHATN-UHFFFAOYSA-N |  | ||
| BGC Download:Ideal Coordinates CCD File | E [auth A] | beta-D-glucopyranose C6 H12 O6 WQZGKKKJIJFFOK-VFUOTHLCSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 76.392 | α = 90 |
| b = 64.379 | β = 98.3 |
| c = 81.003 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Health Research Council (HRC) | New Zealand | 19/397 |